Artificial intelligence has already shaved years off research into protein engineering. Now, for the first time, scientists have synthesized proteins predicted by an AI model in the lab and found them to work just as well as their natural counterparts.
The research used a deep-learning language model for protein engineering called ProGen, which was developed by the company Salesforce AI Research in 2020. ProGen was trained, on 280 million raw protein sequences from publicly available databases of sequenced natural proteins, to generate artificial protein sequences from scratch.
To evaluate if the AI could generate functional artificial protein sequences, the researchers primed the model with 56,000 sequences from five different families of lysozymes, a type of enzyme found in human tears, saliva, and milk, capable of dissolving the cell walls of certain bacteria. The fine-tuned model produced a million sequences, from which 100 were selected to create artificial proteins to test and compare with naturally occurring lysozymes.
The researchers hope that with ProGen generating sequences within milliseconds, it can generate large protein databases that can outstrip naturally occurring libraries.
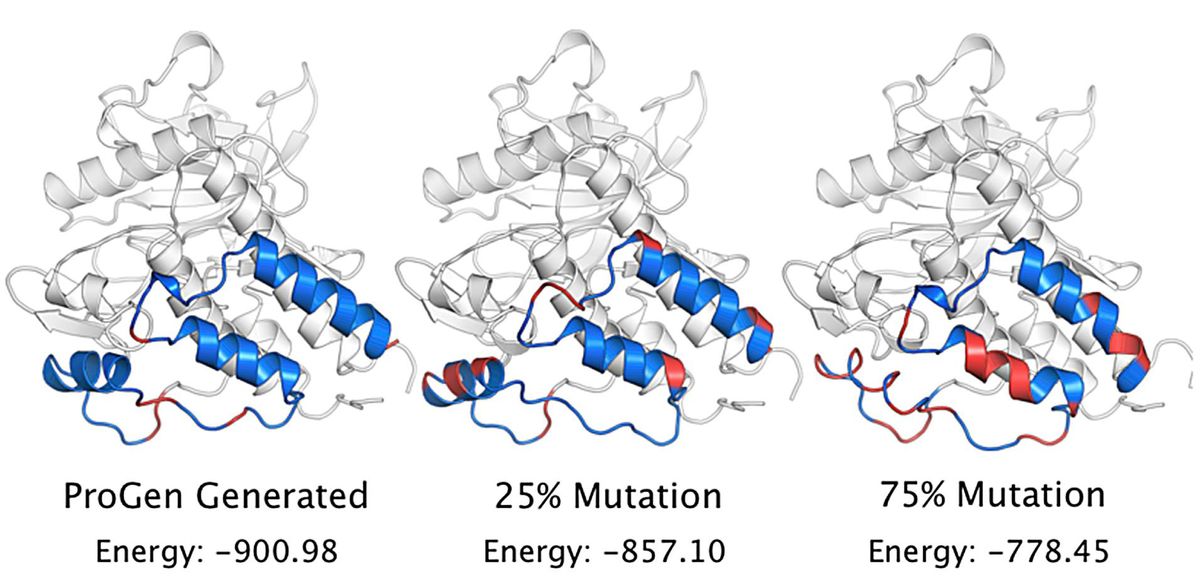
Some 70 percent of the artificial protein sequences worked, too. That, says James Fraser of the University of California, San Francisco’s School of Pharmacy, one of the study’s coauthors, was “not just our top one or two favorites, but actually a statistically meaningful, large number of them.” In fact, Fraser reports, the activity for natural proteins was a little bit lower than for artificial ones. The latter were also active when their resemblance to natural proteins was as low as 31.4 percent.
“What that tells me is that when we use ProGen to generate artificial sequences, those proteins have as good a shot at being active as if we were to choose random natural proteins from the database,” he says, “That, I think, is a big breakthrough.” Researchers, in other words, now have a broader and deeper design palate for protein engineering.
While both physics-based and evolutionary approaches to protein design have worked well so far, Fraser says, these methods have been limited in scope and the chemical catalytic activity the protein displays. This opens up a new way to design proteins with different types of activities in regions of the protein sequence that evolution has not yet explored, he adds.
A key feature of ProGen is that it can be fine-tuned using property tags such as protein family, biological process, or molecular function. “So we can say, Give us [protein sequences] that are, for example, more likely to be thermostable, less likely to interact with other proteins, or potentially better to work with under acidic conditions,” Fraser says. “Having that control, rather than starting from a natural [protein] sequence and trying to coax it to have those properties…is a big dream of protein engineering.”
The researchers hope that with ProGen generating sequences within milliseconds, a large database can be created to expand protein-sequence diversity beyond naturally occurring libraries. This would help find proteins capable of novel catalytic reactions that are related to natural protein activities. For example, says Fraser, “catalyzing a related reaction that might have great attributes for degrading plastic…[or in the] synthesis of a drug…. Being able to go out into sequence space increases the probability of finding that novelty.”
He predicts that the next exciting step for the field will be combining deep-learning language models with other protein engineering approaches to get the best of both worlds—and in the process help researchers find novel activities faster. In the near future, he says, the applications coming out of this research are likely to be about creating new enzymes that could be useful for making small-molecule drugs more cleanly, as well as in the natural process of removing contamination from waste, a.k.a. bioremediation.
Nikhil Naik, director of research at SalesForce, says that their goal was to demonstrate that it is possible to deploy large-language models to the problem of protein design, making use of publicly available protein data. “Now that we have demonstrated that [ProGen] has the capability to generate novel proteins, we have publicly released the models so other people can build on our research.”
Meanwhile, they continue to work on ProGen, addressing limitations and challenges. One of these is that it is a very data-dependent approach. “We have explored incorporating structure-based information to improve sequence design,” Naik says. “We’re also looking into how [to] improve generation capabilities when you don’t have too much data available for a particular protein family or domain.”
The researchers reported their results in the 26 January issue of Nature Biotechnology.
Payal Dhar (she/they) is a freelance journalist on science, technology, and society. They write about AI, cybersecurity, surveillance, space, online communities, games, and any shiny new technology that catches their eye. You can find and DM Payal on Twitter (@payaldhar).