Python has long been one of—if not the—top programming languages in use. Yet while the high-level language’s simplified syntax makes it easy to learn and use, it can be slower compared to lower-level languages such as C or C++.
Researchers from MIT’s Computer Science and Artificial Intelligence Laboratory (CSAIL) hope to change that through Codon, a Python-based compiler that allows users to write Python code that runs as efficiently as a program in C or C++.
“Regular Python compiles to what’s called bytecode, and then that bytecode gets executed in a virtual machine, which is a lot slower,” says Ariya Shajii, an MIT CSAIL graduate student and lead author on a recent paper about Codon presented in February at the 32nd ACM SIGPLAN International Conference on Compiler Construction. “With Codon, we’re doing native compilation, so you’re running the end result directly on your CPU—there’s no intermediate virtual machine or interpreter.”
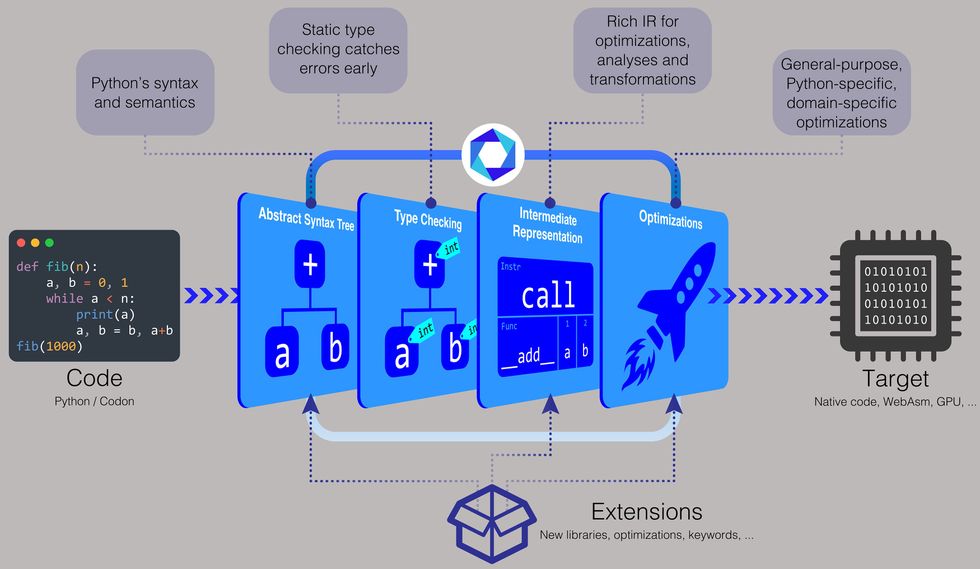
The Python-based compiler comes with pre-built binaries for Linux and macOS, and you can also build from source or build executables. “With Codon, you can just distribute the source code like Python, or you can compile it to a binary,” Shajii says. “If you want to distribute a binary, it will be the same as a language like C++, for example, where you have a Linux binary or a Mac binary.”
To make Codon faster, the team decided to perform type checking during compile time. Type checking involves assigning a data type—such as an integer, string, character, or float, to name a few—to a value. For instance, the number 5 can be assigned as an integer, the letter “c” as a character, the word “hello” as a string, and the decimal number 3.14 as a float.
“In regular Python, it leaves all of the types for runtime,” says Shajii. “With Codon, we do type checking during the compilation process, which lets us avoid all of that expensive type manipulation at runtime.”
MIT professor and CSAIL principal investigator Saman Amarasinghe, who’s also a coauthor on the Codon paper, adds that “if you have a dynamic language [like Python], every time you have some data, you need to keep a lot of additional metadata around it” to determine the type at runtime. Codon does away with this metadata, so “the code is faster and data is much smaller,” he says.
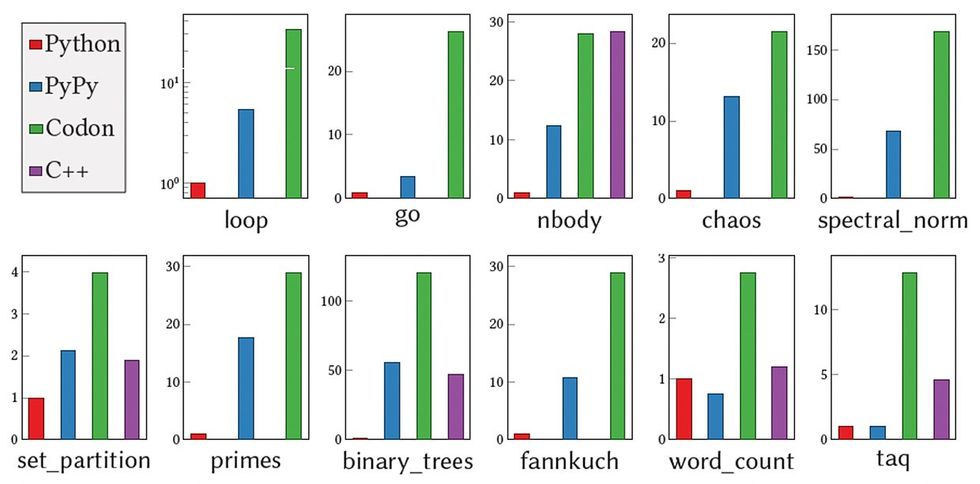
Without any unnecessary data or type checking during runtime, Codon results in zero overhead, according to Shajii. And when it comes to performance, “Codon is typically on par with C++. Versus Python, what we usually see is 10 to 100x improvement,” he says.
But Codon’s approach comes with its trade-offs. “We do this static type checking, and we disallow some of the dynamic features of Python, like changing types at runtime dynamically,” says Shajii. “There are also some Python libraries we haven’t implemented yet.”
Amarasinghe adds that “Python has been battle-tested by numerous people, and Codon hasn’t reached anything like that yet. It needs to run a lot more programs, get a lot more feedback, and harden up more. It will take some time to get to [Python’s] level of hardening.”
Codon was initially designed for use in genomics and bioinformatics. “Data sets are getting really big in these fields, and high-level languages like Python and R are too slow to handle terabytes per set of sequencing data,” says Shajii. “That was the gap we wanted to fill—to give domain experts who are not necessarily computer scientists or programmers by training a way to tackle large data without having to write C or C++ code.”
Aside from genomics, Codon could also be applied to similar applications that process massive data sets, as well as areas such as GPU programming and parallel programming, which the Python-based compiler supports. In fact, Codon is now being used commercially in the bioinformatics, deep learning, and quantitative finance sectors through the startup Exaloop, which Shajii founded to shift Codon from an academic project to an industry application.
To enable Codon to work with these different domains, the team developed a plug-in system. “It’s like an extensible compiler,” Shajii says. “You can write a plug-in for genomics or another domain, and those plug-ins can have new libraries and new compiler optimizations.”
Moreover, organizations can use Codon for both prototyping and developing their apps. “A pattern we see is that people do their prototyping and testing with Python because it’s easy to use, but when push comes to shove, they have to rewrite [their app] or get somebody else to rewrite it in C or C++ to test it on a larger data set,” says Shajii. “With Codon, you can stay with Python and get the best of both worlds.”
In terms of what’s next for Codon, Shajii and his team are currently working on native implementations of widely used Python libraries, as well as library-specific optimizations to get much better performance out of these libraries. They also plan to create a widely requested feature: a WebAssembly back end for Codon to enable running code on a Web browser.
This article appears in the June 2023 print issue as “MIT Makes Python Less Pokey .”
- What Programming Language Skills Do Employers Want? ›
- Top Programming Languages Trends: The Rise of Big Data ›
- C/C++ and Python Top List of Hot Skills for Autonomous Vehicle Engineers ›
- Top Programming Languages 2022 ›
- How Python Swallowed the World - IEEE Spectrum ›
- Python Profiler Links to AI to Improve Code - IEEE Spectrum ›
Rina Diane Caballar is a writer covering tech and its intersections with science, society, and the environment. An IEEE Spectrum contributing editor, she's a former software engineer based in Wellington, New Zealand.



