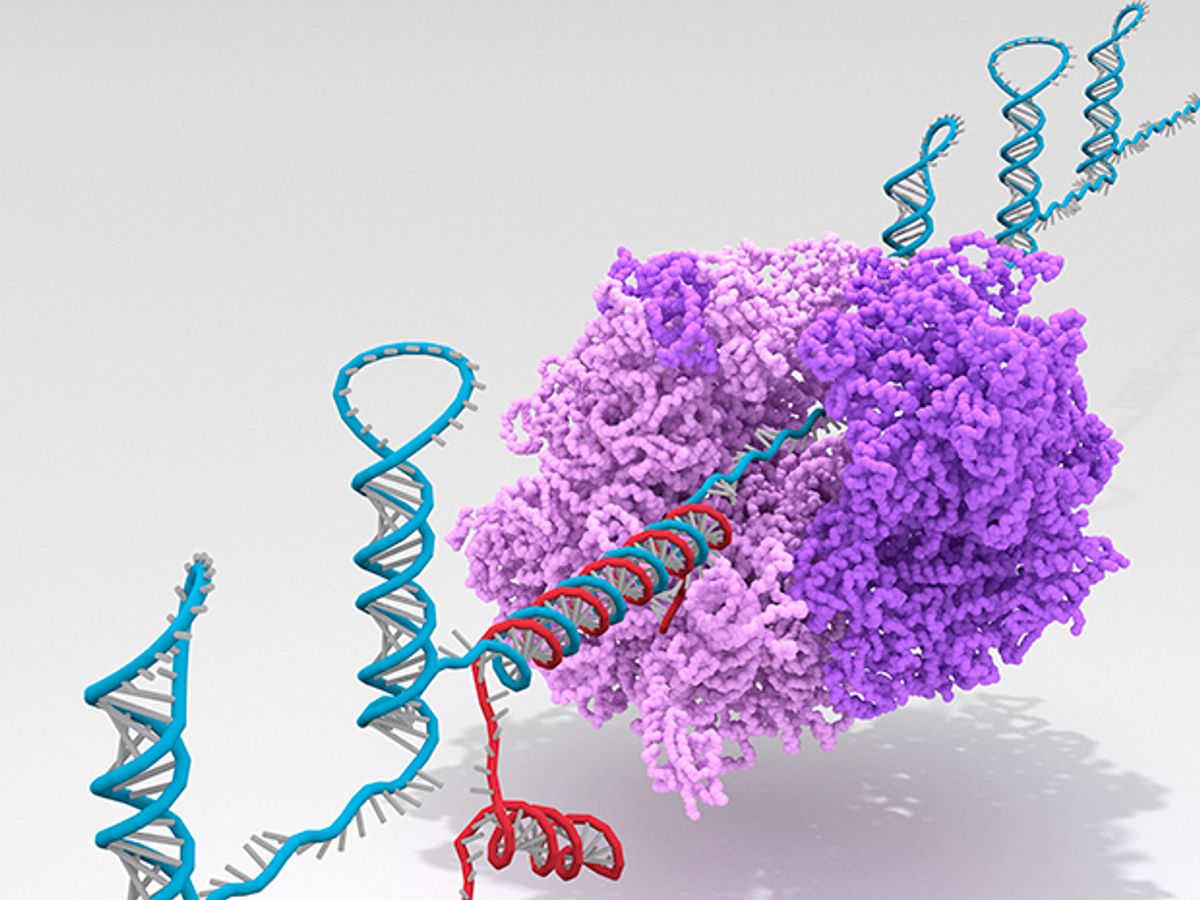
Researchers have developed a biological computer that functions inside living bacterial cells and tells them what to do, according to a report published today in Nature. Composed of ribonucleic acid, or RNA, the new “ribocomputer” can survive in the bacterium E. coli and respond to a dozen inputs, making it the most complex biological computer to date.
“We’ve developed a way to control how cells behave,” says Alexander Green, an engineer at The Biodesign Institute at Arizona State University, who developed the technology with colleagues at Harvard’s Wyss Institute for Biologically Inspired Engineering. The cells go about their normal business, replicating and sensing what’s going on in their environments, “but they’ve also got this layer of computational machinery that we’ve instructed them to synthesize,” he says.
The biological circuit works just like a digital one: It receives an input and makes a logic-based decision, using AND, OR, and NOT operations. But instead of the inputs and outputs being voltage signals, they are the presence or absence of specific chemicals or proteins.
The process begins with the design of a DNA strand that codes for all the logic the system will need. The researchers insert the synthesized DNA into E. coli bacteria as part of a plasmid—a ring of DNA that can replicate as it floats around in the cell.
The DNA serves as a template for the biological computer’s machinery. The cell’s molecular machinery translates the DNA into RNA, essentially copying the DNA code onto a different molecule for use by the cell. RNA links up with a cell’s ribosome and instructs it to produce a protein specified in the RNA’s code.
Here’s where the system behaves like a computer, rather than just a genetically engineered organism: The RNA only does its job when it receives an input that activates it. That’s because the engineered RNA contains codes not just for a protein, but also for logic functions. The logic portions must receive the right inputs in order to activate the RNA in a way that allows the ribosome to use it to produce the circuit’s output—in this case a protein that glows.

Those switches “are kind of the equivalent of your first transistors,” says Green. “They are devices we can build into more complex things.” Green’s team was able to arrange multiple biological transistors, which they call “toehold switches”. Together, the toehold switches form what the researchers called the “gate RNA”, a kind of molecular logic circuit.
The inputs, in Green’s demonstration, were strands of RNA added to the cells after the plasmid containing the “gate RNA” code has already been inserted. These input RNAs paired up with those of the codes of the ribocomputer’s gate RNA. A match between the input RNA and the gate RNA freed the strand so that when it encountered a ribosome, it could tell the cellular machinery to produce the florescence protein. It was easy to see when the system worked—the E. coli lit up.
In their most complex ribocomputer design, they built in enough switches to generate a 12-input circuit made up of 5 AND gates, 5 ORs, and 2 NOTs. That’s really hard to do in a living cell, says Julius Lucks, an RNA engineer at the Center for Synthetic Biology at Northwestern University, who was not involved in the report. “There’s a million ways you can fail at this thing,” he says. In the cell, “everything is interacting with everything else and there are a million ways those interactions could flip the switch on accident.”
Green’s team succeeded in this by using a combination of algorithms that predict how an an RNA molecule folds, along with deep insight into which RNA structures might perform the computations, Lucks says. “It’s the greatest number of inputs in a circuit that a cell has been able to process,” adds Green. “To be able to analyze those signals and make a decision is the big advance here,” he says.
Previous work utilizing DNA and proteins to build biological circuits has seen relatively modest success. The best of those circuits have functioned only in test tubes, rather than in living, replicating cells, or they weren’t able to process as much information in such a compact space as the ribocomputer. For example, Microsoft researchers this week reported a method for DNA computing using an origami-style structure. The design enabled faster computation than other such DNA circuits, but it wouldn’t work inside a living cell.
Christopher Voigt at MIT has created software that automates the design of DNA circuits for living cells, and last year demonstrated its capabilities. But those designs couldn’t handle as many inputs as the new ribocomputer technology and required more DNA to encode.
Green says he hopes DNA- and RNA-based computing will pick up speed now. “We’re like 70 years behind where things are in electrical engineering,” he says. (The first transistor was invented in 1947.) “Once we get the infrastructure for building biological circuits, hopefully we’ll have similar growth,” he says.
Green says he envisions biological circuits being used in diagnostic devices that detect viruses or in cellular devices programmed to break down environmental toxins.
Or it could even be used to prevent the spread of disease. “You could upload a biological circuit into the cells in the body to protect against viruses or prevent cancer from developing,” he says. “That is, the diagnostic would be part of your own cells.” If the body acquires cancerous or viral materials, cells could be programmed to shut down or synthesize a drug or summon the immune system into action, instead of allowing the disease to spread.
This post was corrected on 26 July to fix the illustration caption.
Emily Waltz is the power and energy editor at Spectrum. Prior to joining the staff in January 2024, Emily spent 18 years as a freelance journalist covering biotechnology, primarily for the Nature research journals and Spectrum. Her work has also appeared in Scientific American, Discover, Outside, and the New York Times. Emily has a master's degree from Columbia University Graduate School of Journalism and an undergraduate degree from Vanderbilt University. With every word she writes, Emily strives to say something true and useful. She posts on Twitter/X @EmWaltz and her portfolio can be found on her website.