Following DNA Down the Nanopore Rabbit Hole
If we ever want to make a DNA-read-head, we need to understand how DNA moves through tight spaces--new research is doing just that.
DNA is a linear information storage medium, like a magnetic tape, but written with a very limited character set—just A, C, G, and T.
It would obviously be much quicker, cheaper, and more reliable if we had a DNA-read-head, a device we could just run the DNA-tape over to detect the sequence directly, in a single pass. Back in 1995, George Church and colleagues figured out one approach: it might be possible to use a low intensity electric current to pull long strands of DNA through nanometer-scale pores in a membrane and measure the electric field variations of the four nucleic acids—A, C, G, T—as they passed through.
Scientists are working on it, but we’re not there yet. Key questions remain unanswered. One of them is fundamental: How does DNA move through a pore? Does it slide through end-on? Or does the pore grab it somewhere in between, bend it double, and suck it through doubled over (like a strand of spaghetti slurped up from the middle). Every cell in your body, after all, carries about two or three meters of DNA. Even a mere fragment of 50,000 base pairs is about 16.5 micrometers long—thousands of times the diameter of a nanopore. If mere chance determines the orientation, one would expect that almost all DNA should pass through a nanopore doubled over.
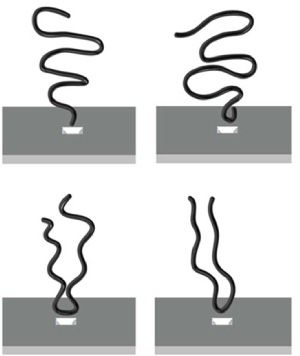
Fortunately for the future of bedside genomics, that doesn’t seem to be the case. Researchers at Brown University have developed an elegant method for determining exactly how a DNA molecules passes through a nanopore. They can see if it slips through end-on or goes through doubled over…and, if it does double over, they can tell where along its length it folds.
The researchers found that DNA passes through the pore smoothly, end-on, about 25% of the time—far more than most current models would predict. This surprisingly high proportion—indicating, they say that the orientation is a function of “the configurational entropy of the approaching polymer”—bodes well for developing a nanopore-based DNA direct reader.
Figures: Derek Stein/Brown University
Douglas McCormick is a freelance science writer and recovering entrepreneur. He has been chief editor of Nature Biotechnology, Pharmaceutical Technology, and Biotechniques.