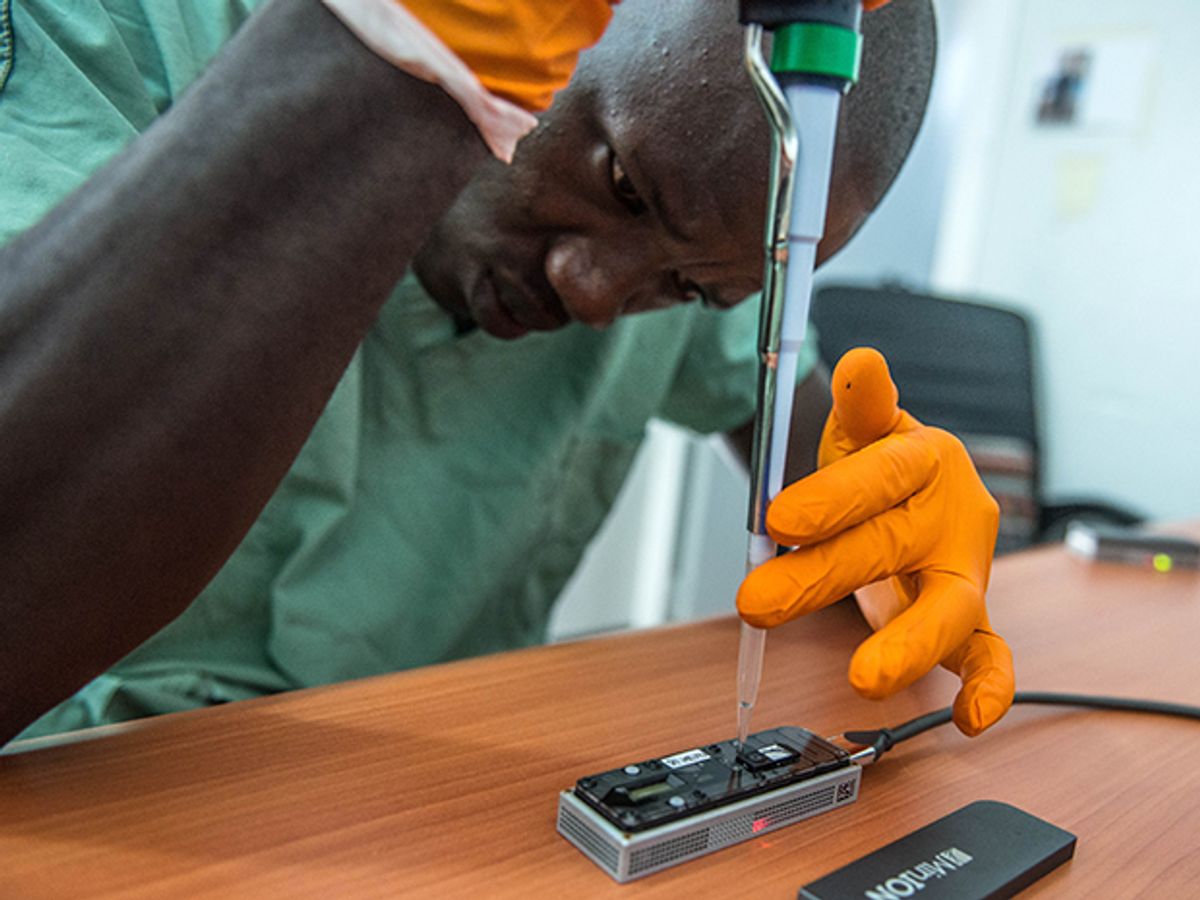
Using a genome sequencer smaller than a stapler, geneticists have demonstrated the role they can play in combating outbreaks of infectious disease. An eight-month experiment in Guinea during the tail end of the Ebola outbreak, described today in a the journal Nature, showed the potential of a genome sequencing technology that can be packed inside a suitcase and deployed in rural outposts.
With the Zika virus outbreak gaining momentum in the Americas, the Ebola experiment may offer useful lessons. “Having genome data is becoming part of the fundamental response to an outbreak,” says lead researcher Nick Loman, a geneticist at the University of Birmingham. By studying the genetic material of the virus across many patients, researchers can look for telltale mutations that reveal the paths of transmission. And if those routes are discovered quickly enough, public health officials could make decisions to change the course of the epidemic.
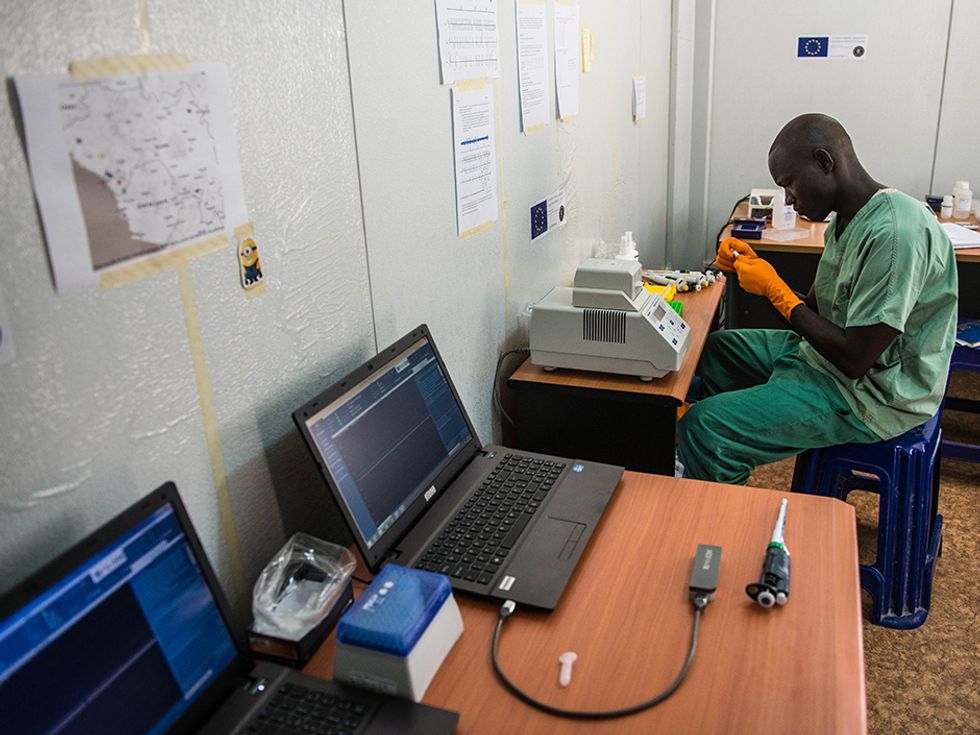
In the Ebola study, Loman and his colleagues used the MinIon sequencer from Oxford Nanopore Technologies. This diminutive device plugs into a laptop via a standard USB port, through which it both transmits data and draws power. The researchers packed three of these devices, three laptops, and all the other equipment and reagents they’d need into the suitcases shown in the photo below—“the rucksack on top just contains pants and socks,” says Loman. “This was the first time we could put an entire genome sequencing laboratory into aircraft-hold luggage.”

One of Loman’s graduate students, Joshua Quick, brought the gear to Guinea in March 2015, and worked with the European Mobile Lab project to get set up quickly. Within two days, the team was sequencing samples of the Ebola virus drawn from local patients. The Ebola virus’s genome is composed of about 19,000 “letters” of genetic material called RNA; sequencing is the task of putting all those letters in the proper order. Then researchers could compare the genome sequences of different patients, looking for mutations that have reordered those letters along some stretches of the RNA strand. If patients have a virus withthe same mutations, they were likely part of the same person-to-person chain of transmission.
The Guinea team regularly gave their results to public health officials to help them draw conclusions about the virus’s spread. “This started as a test run, and turned into real-time surveillance that was feeding into the epidemiology,” Loman says. For the most part, the results just confirmed known chains of transmission. But as the outbreak drew to a close in late 2015, the team provided useful intelligence on final flare-ups. “Occasionally a case comes out of the blue, and the question is, where did that case come from?” Loman says. “With genome sequencing you can say, it’s linked to this cluster.” Then public health officials can take the necessary steps to isolate the area.
During the West Africa Ebola outbreak, researchers tried out a mobile genetics lab that allowed them to study the Ebola virus’s genome on the ground in Guinea.
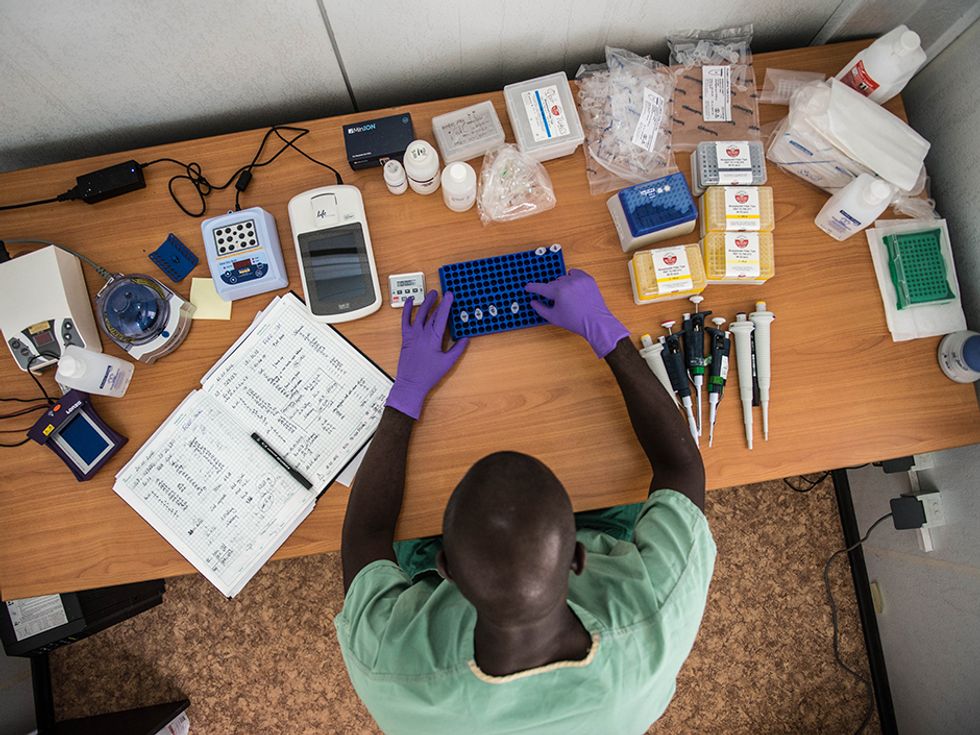
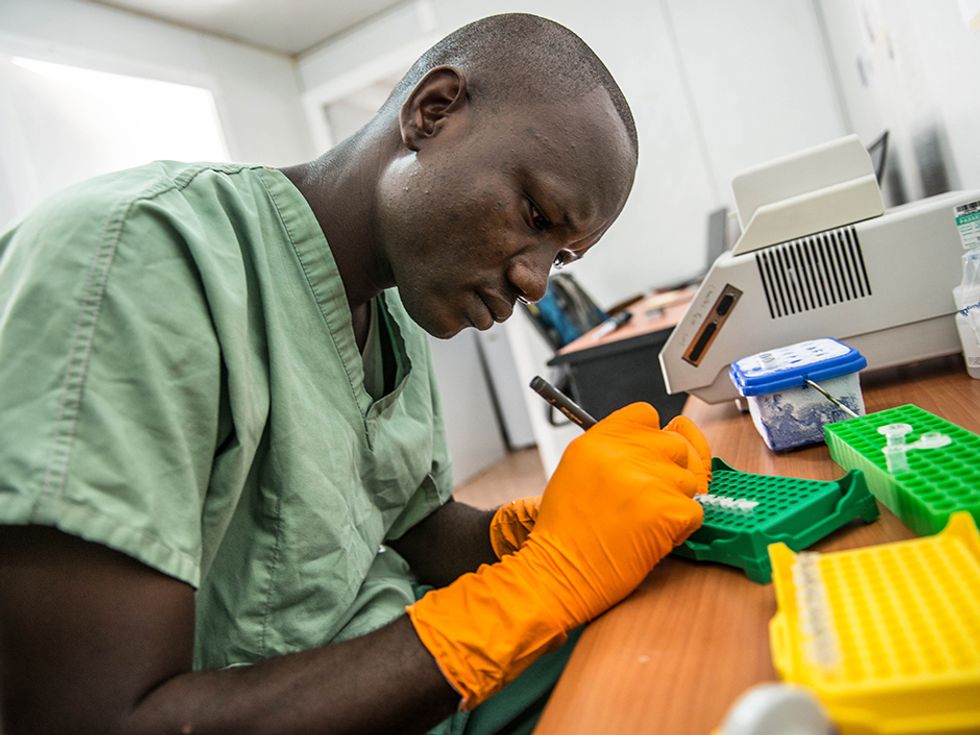
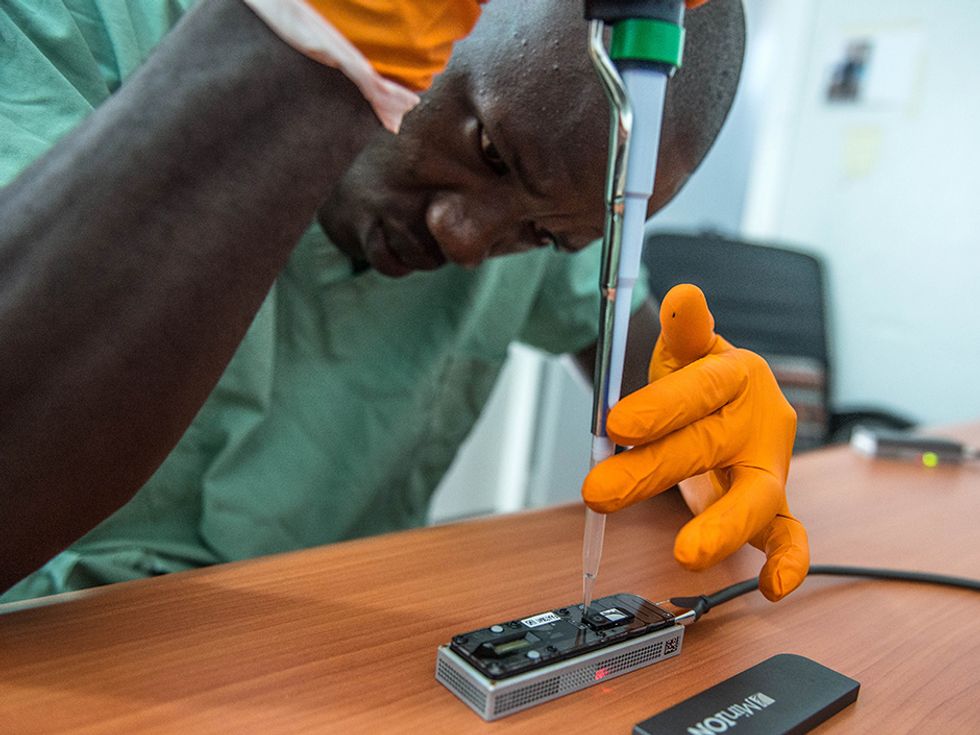




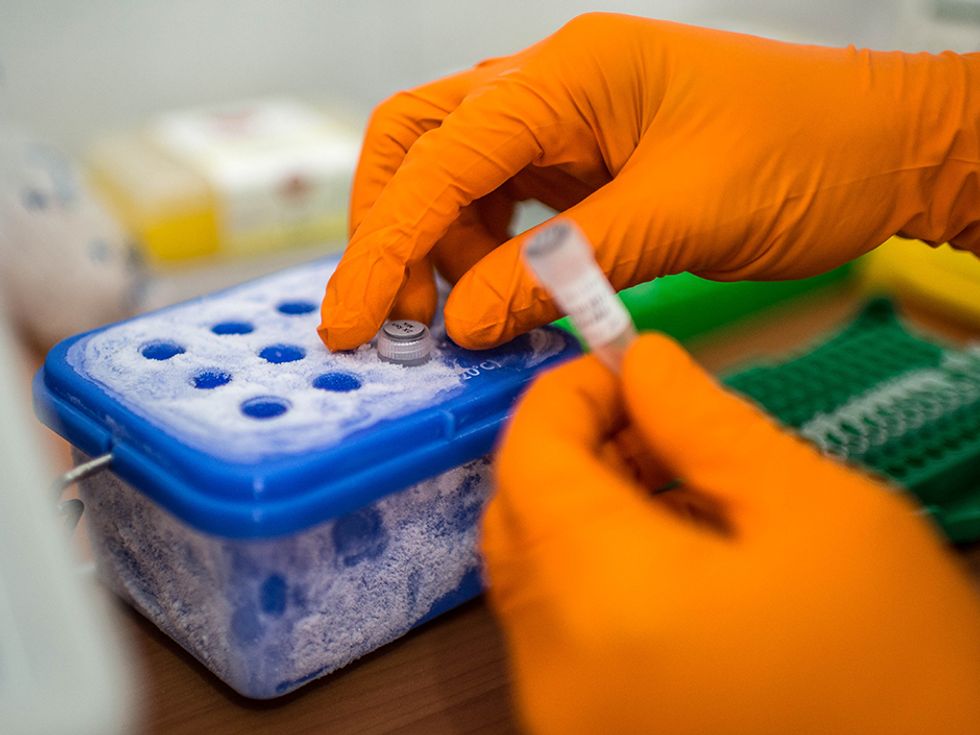

The MinIon sequencer uses a fairly new type of sequencing technology, quite different that the methods used by the hulking machines that dominate the market. Inside the MinIon sequencer, an electric current is sent through a membrane poked through with tiny nanopores. When strands of DNA or RNA pass through those pores they alter the current, with the different types of letters in the strand altering the current by different amounts. Recording the sequence of electric signals in the device thus reveals the sequence of letters in the DNA or RNA. The MinIon isn’t yet commercially available, but Oxford Nanopore has invited many researchers to try it out by paying an access fee of $1000 and joining the MinIon “community.”
This high-tech device worked just fine in the simple mobile labs in Guinea, Loman says. The big problem: “Power and internet,” he says. Power cuts took some of their instruments offline until the team invested in a backup power source, and agonizingly slow upload speeds limited their ability to crunch the data.
Frontline workers in South America who are now trying to contain the Zika virus’s spread have somewhat different needs than did the Ebola responders. While Ebola is easily transmitted from person to person, Zika virus is typically spread when a mosquito bites an infected person and then goes on to bite someone else. But public health workers will still need to figure out how the virus is spreading around the world in this age of frequent air travel, and genome sequencing will help with that. Genomic information can also help researchers devise diagnostics and develop vaccines, Loman says. “I like to think that people will pick up our work, and use it with this outbreak,” he says.
Eliza Strickland is a senior editor at IEEE Spectrum, where she covers AI, biomedical engineering, and other topics. She holds a master’s degree in journalism from Columbia University.