The Gene Machine and Me
Ion Torrent’s chip-based genome sequencer is cheap, fast, and poised to revolutionize medicine

It’s a fresh April morning in 2012 when I head to Connecticut to see a man about a genome. Not just any genome, but my own.
I want to learn my own biological secrets. I want to get a look at the unique DNA sequence that defines my physical quirks, characteristics, and traits, including my nearsighted blue eyes, my freckles, my type O-positive blood, and possibly some lurking predisposition to disease that will kill me in the end. So I’m not going to see just any man, but the mad scientist of genomics himself, the arrogant upstart of biotechnology, an inventor and entrepreneur who has upended the business of genetic sequencing once before—and now appears to be doing it again.
Jonathan Rothberg, founder and CEO of the company Ion Torrent, believes his cheap and fast sequencing machines will revolutionize medical practice within the decade. Indeed, he says, the revolution has already begun. In some hospitals, cancer patients can already have part of their genomes checked before their physicians decide on treatment. Newborns with life-threatening problems can have their genomes scanned to give doctors insight into what’s wrong—hopefully, before it’s too late.
Soon, Rothberg says, everybody will be sequenced—probably as infants—and will be able to make diet, lifestyle, and medical choices based on specific information, rather than on hunches about vulnerabilities. Knowledge of a person’s genome will allow specialists to customize medical treatments and drugs for that patient, to maximize effectiveness while minimizing side effects. A routine checkup could start with the doctor checking for “updates” to a patient’s genomic file; if medical research has turned up new data about the patient’s particular set of genes, the doctor would get an alert. “Our goal,” Rothberg declares, “is to really transform medicine.”
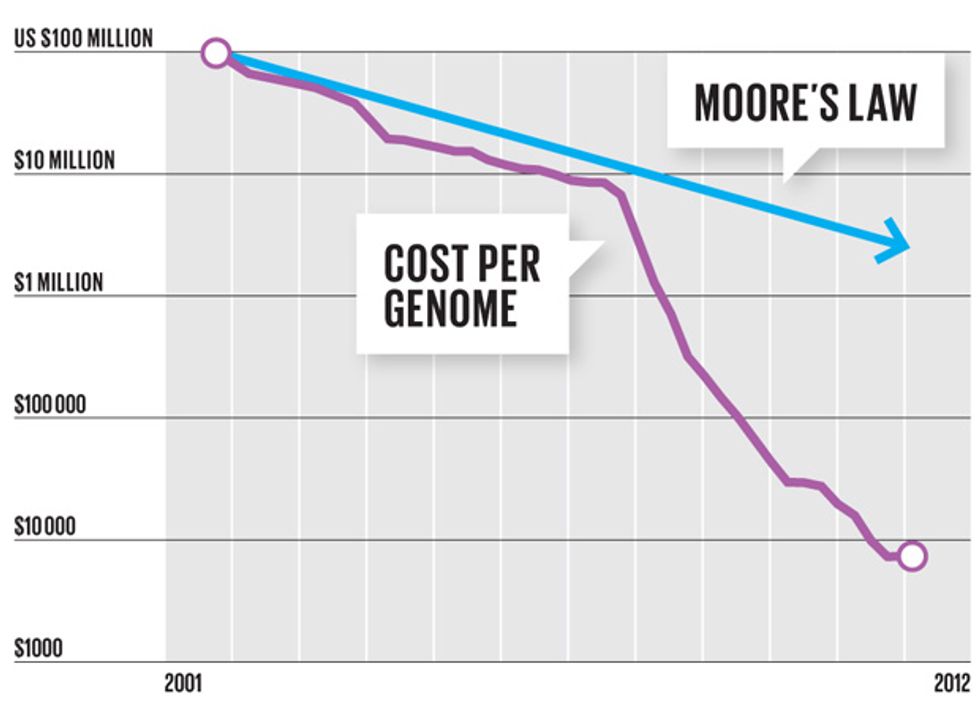
To decode the first human genome, a milestone completed 10 years ago, armies of researchers labored for more than a decade and spent more than US $3 billion. Working with Rothberg’s newest machine, a technician will soon be able to decipher a human genome in a few hours, and at the bargain-basement price of $1000.
My personal quest for genetic enlightenment begins with a visit to Ion Torrent’s black glass headquarters in a Guilford, Conn., office park. Ion Torrent isn’t really Rothberg’s baby anymore—the company was bought by biotech giant Life Technologies Corp. in 2010 for $725 million—but he still strides through the facility with a proprietary air. He beams as we tour labs where biochemists and electrical engineers work side by side. The windows, pressed into service as extra whiteboards, are covered with scribbled sketches of molecular diagrams and circuit diagrams alike.

Several other companies are also racing toward the $1000-genome goal. But Rothberg was the first to tap into the accumulated expertise of the computer industry with a method he calls “semiconductor sequencing.” Inspired by the cover of a computer magazine in a hospital waiting room, he figured out how to use an integrated circuit to record how a DNA molecule is built out of chemical components, and how to turn that chemical information into a digital readout.
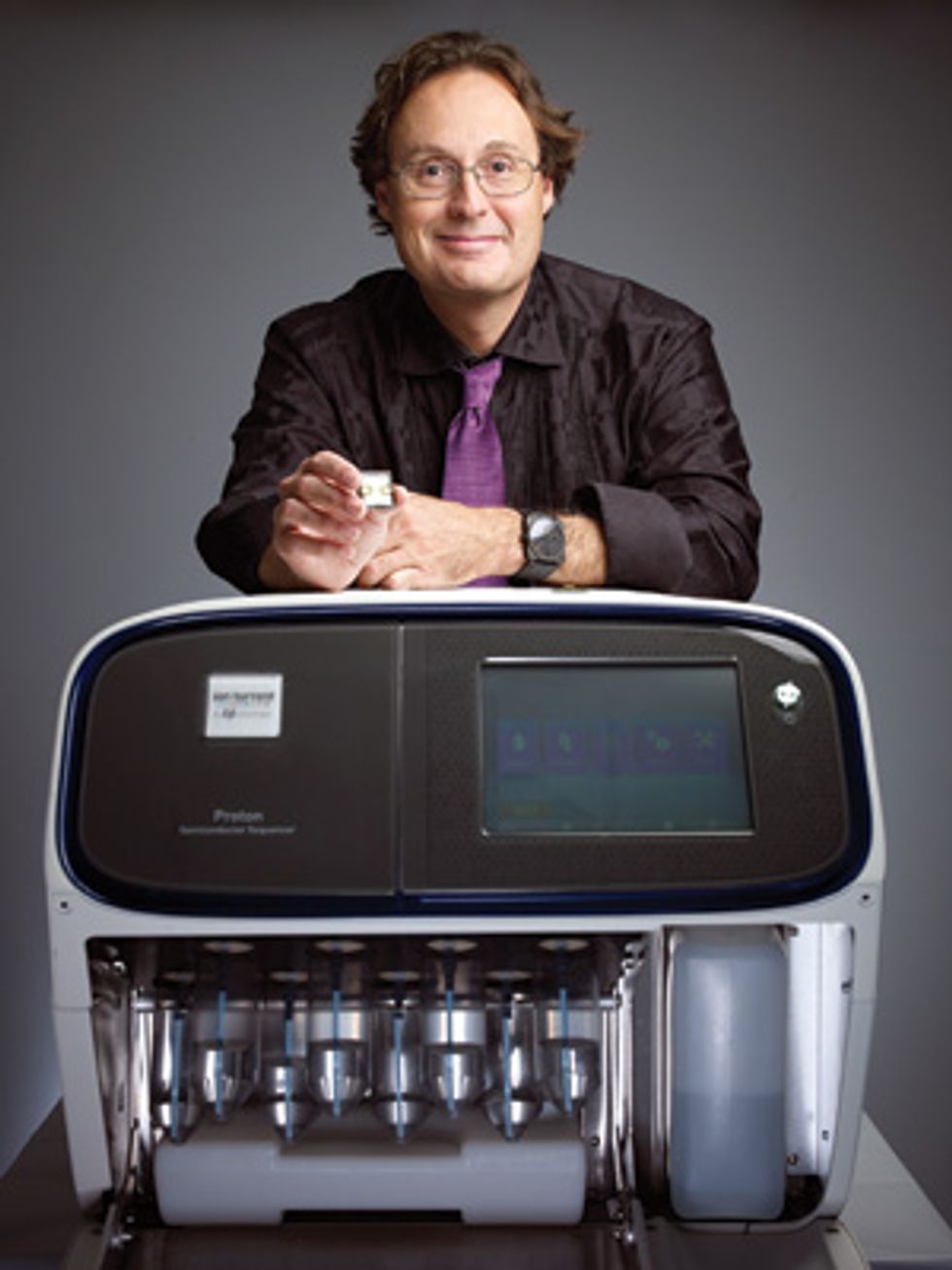
Rothberg’s newest product is a sleek machine called the Ion Proton System, which is just a little bigger than a fancy office printer. With just three disposable chips and a few hours of time, that machine can make sense of the genome of a human being, identifying each of the 3 billion molecules that make up a person’s DNA. Ion Torrent’s current top-of-the-line production chip isn’t much by semiconductor-industry standards—it’s got a measly 400 million transistors. But by tapping into the existing supply chain for microchip manufacturing, Rothberg has already improved his chips nearly a thousandfold, and he’s not finished. Ion Torrent plans to ride Moore’s Law as far as it will go.
Rothberg wants to be very clear about what all this means to medicine. “Look at the things that have added to life expectancy,” he says during our first meeting. “It’s clean water—that was the single biggest health improvement. Next came the antibiotics, next probably imaging, whether it’s X-ray or MRI or CAT scan.” He flashes a cocky smile. “I do believe that genome sequencing will have as profound an effect on medicine as clean water, antibiotics, and imaging.”
I’ve come to Connecticut to learn about this hot new machine that might, in the words of one of the company’s taglines, provide “sequencing for all.” But I don’t just want to hear about how it works; I want to try it out for myself. I know that having my genome sequenced will probably reveal something unusual about my genetic makeup. Quite possibly, that something will be disconcerting.
“We are here,” said U.S. president Bill Clinton, standing in the White House in June 2000, “to celebrate the completion of the first survey of the entire human genome. Without a doubt, this is the most important, most wondrous map ever produced by humankind.” The map showed the placement of the 3 billion “letters” that spell out the human genome—all the genetic material contained in Homo sapiens.
Forgotten the genetics you learned in high school biology? Here’s a refresher. Every cell in your body contains deoxyribonucleic acid, or DNA. That DNA is built from four molecules called nucleotides and identified by the letters A, G, C, and T, which stand for adenine, guanine, cytosine, and thymine. Think of DNA as a pair of chains, each shaped into a helix. The nucleotides are the links in the chain, and in DNA they pair up: One half of each pair is attached to one helix; the other half is attached to the other one. A always pairs with T, G always with C.
Genes are specific groups of letters on the long DNA chain. Each gene encodes the instructions for producing one of tens of thousands of proteins that conduct the business of life within our bodies. They include the hemoglobin that carries oxygen, the antibodies that fight infections, and the enzymes that permit the digestion of food, the contraction of muscles, and countless processes in the brain. By defining which proteins are produced and whether they’re normal or irregular, the genome exerts enormous control over how we look, how we feel, and how we act—not to mention how and when we get sick.
Understanding the genome has long been a fundamental goal, and the federally funded Human Genome Project (along with a private company, Celera Genomics) completed a “rough draft” in June of 2000 and a final version three years later. The researchers analyzed genetic material from a handful of people to arrive at the sequence of letters that defines a typical human being. Every person’s genome differs from that reference genome at a relatively small number of places in the long sequence of letters, but those tiny differences are enough to make us individuals.
Since the Human Genome Project launched in 1990, sequencing technology has advanced at a breathtaking pace, which genetic researchers speak about with a mix of awe and bitterness. “For my Ph.D., I spent six years and I sequenced 9000 letters,” Rothberg says. He looks me in the eye and says it again, in a pained voice. “Six years. For 9000 letters.” Rothberg was a Ph.D. student in the late 1980s, when the state-of-the-art method required researchers to laboriously analyze one DNA fragment at a time. In the 2000s lab techs rejoiced over the introduction of “massively parallel” machines able to analyze many fragments of DNA at once.
The first of those machines emerged from Rothberg’s prior company, 454 Life Sciences. With a showman’s flourish, Rothberg proved that machine’s utility in 2007 by sequencing the entire genome of James Watson, one of the genetic pioneers who discovered the double-helix shape of DNA in 1953. The effort that Rothberg dubbed “Project Jim” took two months and was hailed as a breakthrough, for Watson’s was the first individual genome to be sequenced, and it revealed the extent to which each unique human varies from the average. Watson, who has never been described as shy, promptly put his genome on the Internet.
The Competition
Other biotech companies and start‑ups are also striving to dominate the genome-sequencing market.
Illumina: HiSeq 2500
Illumina’s expensive but reliable machines are the market leaders and can be found in hospitals and labs worldwide. The company is focused on speeding up whole-genome sequencing.Pacific Biosciences: PacBioRS
This machine, released in 2011,can sequence a human genome using single molecules of DNA. The technique provides highly accurate results.Complete Genomics
Complete Genomics doesn’t sell its machines to researchers. Instead it offers them in-house genome sequencing and analysis packages.Oxford Nanopore: GridIon
This start-up hasn’t released a commercial machine yet, but there’s considerable excitement about its single-molecule sequencing technology.
As the technology made genetic scans faster and cheaper, doctors found ways to use them. Physicians have begun ordering scans for patients on diagnostic odysseys—chronically sick patients who have gone in for test after test without finding the cause of their illnesses. Cancer hospitals have started sequencing both their patients’ regular genomes and the mutated versions found in tumors; this allows doctors to study genetic differences that encourage a tumor’s growth and lets them look for drug targets. And massive research efforts are under way around the world. By sequencing thousands of people with autism, diabetes, or any other number of complex conditions, researchers hope to identify the specific genetic variations that are common within these groups.
Last March, a study from the insurance company UnitedHealth Group predicted that spending on genetic tests will skyrocket from about $5 billion in 2010 to between $15 billion and $25 billion in 2021. That market includes not only medical-grade scans, which a physician orders and interprets for a patient, but also tests that are sold directly to consumers by companies such as 23andMe, in Mountain View, Calif. These companies market their tests to the curious and the health-conscious and offer a very limited set of genetic results, most of which are interesting but not medically significant. Customers learn about genealogy, a few risk factors of questionable importance, and oddities like whether they’re able to smell asparagus in their own urine.
I tell Rothberg that I want medically relevant results, and he puts me in touch with the pioneering Human Genome Sequencing Center at Baylor College of Medicine, in Houston, which collaborated on the Watson project. Baylor’s researchers are working with Ion Torrent to validate its new Proton sequencing machine, and they agree to give me early access to the technology and to run my DNA through one of their Proton machines. I would be Baylor’s first merely curious patient, but surely not its last.
In late August, I find myself cozily ensconced in a Baylor conference room with the geneticist James Lupski, a professor in the medical school. The silver-haired physician had the task of telling James Watson his genome results in 2007. “I had to be the one to say, ‘Well, Jim, we don’t know what the hell your DNA means, because you’re the first one to be sequenced,’ ” Lupski says with a crackling laugh.
Since then, Baylor researchers have been pushing to make genome scans useful. “There was the question, could we find things that were important for medical management?” Lupski says. “Was the signal above the noise?” The noise, he explains, is the thousands of genetic variants found in each individual, because “everybody truly is unique.” In 2011, Lupski and his colleagues answered that question with 14-year-old twins who suffered from a palsy-like movement disorder. Whole-genome sequencing revealed not only the genetic mutation responsible, it also suggested a new drug regimen that worked remarkably well to control the twins’ symptoms.
But the twins are still the exception. Baylor opened a commercial lab in October 2011 to provide sequencing services for doctors grappling with tough cases—patients who are on “medical mystery tours,” as Lupski puts it. The lab has a 30 percent diagnostic success rate, which means doctors can pinpoint, in almost one-third of the cases, the mutations in a patient’s DNA that are causing symptoms. That rate may seem pretty low. But doctors consider it a remarkable achievement, given the sprawling complexity of any human’s DNA and how little is now conclusively known about how our genes function. As Lupski puts it, “in this branch of medicine, we admit our ignorance a lot more.”
In the Baylor conference room, Lupski explains what his team will do for me: They will sequence only my exome, the portions of the genome that contain genes. While the 30-million-letter exome makes up just 1 percent of the genome, changes to those portions account for the vast majority of genetic diseases. The rest of the letters that make up the genome, historically dismissed as “junk DNA,” play some role in human health, but the details aren’t yet clear. Baylor’s lab currently specializes in exome scans but may eventually switch to whole genome scans once medical understanding advances.
As we go over consent forms, Lupski tells me about all the terrible, horrible things this process could reveal about me. I could find out that I have genetic mutations that put me at high risk of developing a deadly disease like Parkinson’s or cancer. Anything I learn, Lupski intones gravely, will have implications not just for me but also for my immediate family: my parents, my sister, and her children. I might also find out that I’m a carrier of some recessive genetic ailment, meaning that if my husband and I choose to have children and he’s a carrier too, our kids could end up with that disorder. If there’s bad news, do I want to know it?
I do. I’ve already talked the project over with my family. My parents worried at first that I’d discover something that would be hard to live with, but they ultimately gave their blessing. My sister was fine with it, too. “I’m not afraid of information,” she said. So I tell Lupski not to hold anything back. I hope I’ll get early warnings about problems I may encounter later in life and suggestions on screening regimens or lifestyle changes that can protect me.
I’m a trailblazer: Almost everyone tested at Baylor before me had one or more known serious diseases. I do not—as far as I know. But “what you find is that a lot of us have more susceptibilities to disease than anyone suspected,” Lupski tells me. “Many of us are walking around with all kinds of things.” As sequencing keeps getting cheaper, it will eventually become standard practice to screen healthy individuals for their vulnerabilities, Lupski believes. Just as physicians today take family medical histories from their new patients, they may soon order exome scans as well.
Lupski agrees to give me everything judged medically relevant, but no more: “We’re not going to tell you what your genes say about the color of your hair or your eyes.” No problem. I already know what color they are. I sign the consent forms and stick out my arm to let a doctor draw my blood.
Jonathan Rothberg is big on epiphanies. Right now, in a meeting room at Ion Torrent, he’s telling me about the biggest one of his life, which occurred in a hospital waiting room on 2 July 1999.
Rothberg was then the CEO of a drug discovery company he had founded to trawl through the genetic data being produced by the Human Genome Project. But then, “on July 2 my son Noah was born,” Rothberg says, “and he was having difficulty breathing. No matter how interested I was in this map that bound us all as humans, I really only cared about his genome.” Understanding the genome in the abstract was all well and good, but a scan of Noah’s personal genome could help doctors understand what was wrong—and possibly help them fix it.
While Rothberg paced in the waiting room, his gaze fell on the cover of a computer magazine, InfoWorld. It trumpeted the release of the Pentium III chip, which incorporated 9.5 million transistors. The drive to miniaturize semiconductors had produced the personal computer revolution, Rothberg thought, so maybe shrinking down sequencing technology could usher in a personal genetics revolution.
Noah’s breathing problems passed quickly, and the parents took home their burbling infant. But Rothberg’s revelation obsessed him, and he spent his two-week paternity leave roughing out the idea for a new sequencing machine. “I got in trouble for it,” he said with a laugh. Those sketches led him to found his next company, 454 Life Sciences, which in 2005 introduced the world’s first next-generation sequencing machine.
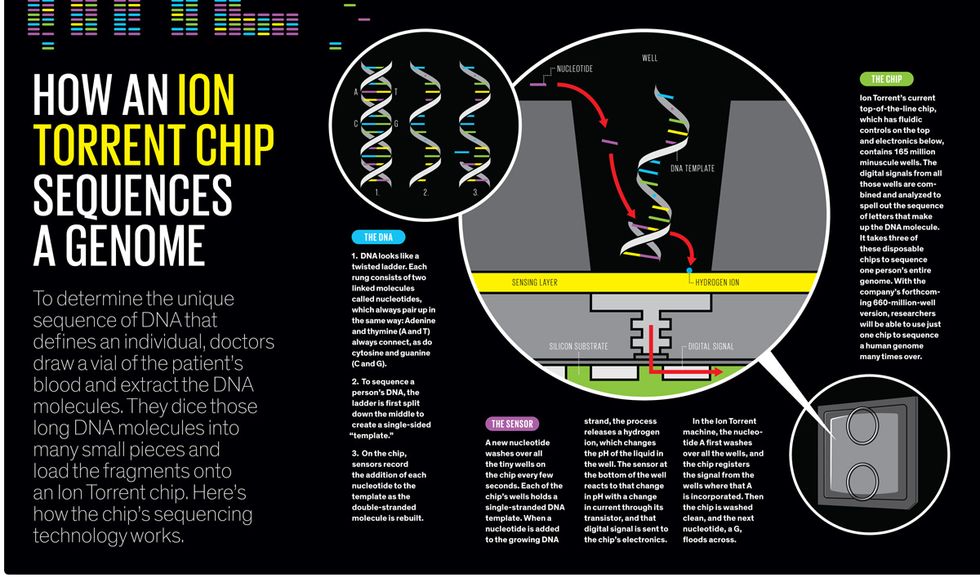
To understand how it worked, recall the DNA molecule. It looks like a twisting ladder, with rungs formed by pairs of As and Ts and pairs of Cs and Gs. This structure is fundamental to the mechanism by which DNA copies itself every time a cell divides. To reproduce itself, the double-stranded helix “unzips” down the middle, making each single strand a template; each A attracts a new T, each C latches onto a G, and so on, until there are two complete twisted ladders instead of one.
Next-gen machines typically rely on a method called sequencing by synthesis, in which a DNA molecule is broken up into many pieces of single-stranded templates. These templates are placed in thousands or millions of minuscule wells on a substrate inside the machine. Then the series of four nucleotides wash over the wells, one at a time, while the machine records which templates add the A, then the G, and so on, through millions of washes. In Rothberg’s process, a firefly enzyme was attached to each nucleotide and produced a flash of light when it was added to the template. A camera inside his 454 Life Sciences machine took a photo with each wash to record the flashes.
While competitors scrambled to catch up, Rothberg promoted his machine with flair. He sequenced Watson’s genome and he collaborated with German researchers to sequence a Neanderthal’s DNA, using genetic material extracted from bone samples. But in 2007 the company was purchased by the biotech giant Roche Diagnostics, leaving Rothberg rich but jobless. He had to start again, and he needed a new idea.
That idea emerged from a fundamental problem that 454 and its competitors had experienced: Inventing a new sequencing technology had also meant creating a new supply chain. The companies all had to establish manufacturing processes for their machines and components, including the substrates, chemicals, and sophisticated optical instruments. “Everybody ran into trouble,” Rothberg said. His next company, Rothberg decided, wouldn’t just take inspiration from the semiconductor industry, it would also adopt its entire supply chain. He set out to design a semiconductor chip that could sequence DNA and be manufactured in a standard microchip foundry.
Rothberg found his solution, surprisingly, in a technology that’s been around for four decades: the ion-sensitive field-effect transistor (ISFET). The electric current flowing through an ISFET submerged in a solution varies depending on the pH of that solution. Rothberg knew that adding a nucleotide to a growing DNA strand naturally releases a hydrogen ion, which makes the solution more acidic. It should be possible, he reasoned, to put many ISFETs in an array to detect the minute pH changes as As and Gs and so forth were added to a DNA strand. “This would be a sensor array that would literally see chemistry,” Rothberg says with a proud grin.
With his new company, Ion Torrent, he built a machine that continued to rely on a plate covered with tiny wells. But instead of indicating the addition of nucleotides with flashes of light, the machine registered them by using an array of ISFETs to detect tiny changes in pH. The first chip he made was a dinky thing with an array of just 256 000 ISFET sensors, all aligned with wells containing pieces of DNA template. When the first nucleotide—say, an A—washed over the chip, all those template pieces that needed an A to complete the first rung of the ladder incorporated it. The ISFET sensors in those wells reacted to the change in pH, and their change in current was registered as digital information.
Rothberg’s technology improved quickly: His second experimental chip had 1 million ISFET sensors, and he has kept on iterating. Increasing the number of sensors and wells means that more template pieces can be analyzed in parallel during a single run of the machine, making sequencing faster and cheaper. Ion Torrent’s forthcoming chip, expected in mid-2013, will have 660 million wells, each with microfluidics on top and an ISFET sensor below.
In our conversations, Rothberg said his technology has improved so rapidly because he’s “catching up with the accumulated Moore’s Law,” the five-decade-old dictum stating that the number of transistors on leading-edge ICs will double about every two years. To drive his point home, in July 2011 he used his machine to sequence the genome of none other than Gordon Moore, author of the eponymous law. It took 1000 chips to decode Moore’s genome then. With Rothberg’s next chip, it will take one.
On a tour of the Guilford labs, Rothberg pauses in front of his latest machine, the Ion Proton. “We’ll have chips that have 2 billion and 5 billion sensors,” he declares. “These machines are fully capable of enabling a $1000 genome. Then as we replace these disposable chips at higher and higher densities, we’ll be able to drop that cost even further.” Most of the cost comes from the single-use chip, which is tossed after a machine’s run; the chemicals used are cheap.
Rothberg is convinced that he’s a visionary. In general, objective observers agree. “When I first heard that Rothberg had developed a sequencing platform essentially by measuring pH, I was astounded, I was flabbergasted,” says Kevin Davies, author of the recent book The $1000 Genome (Free Press, 2010). Davies praises Rothberg for his ability to knit together ideas from such disparate industries as chemistry, engineering, and software. “He pulls the threads together and makes them work,” Davies adds. “That’s his brilliance.” However, Davies notes that there’s plenty of excitement about other emerging technologies, including sequencing techniques that require only a single molecule of DNA to work. Several biotech companies, including Oxford Nanopore Technologies, in England, and Genia Corp., in California, are racing to commercialize those.
Robert Darnell, president of the New York Genome Center, says that Ion Torrent’s technology is fascinating, but he stops short of calling it the market leader. The New York center, just established, will provide genome-sequencing services to New York’s biggest medical and research institutions, and Darnell’s team is therefore buying a lot of machines. After a “careful assessment,” Darnell says, his organization decided to purchase 25 sequencing machines from Illumina, a biotech concern in San Diego. But the genome center will also try out newer machines, including Ion Torrent’s Proton. The situation is changing so fast, Darnell said, that today’s market leader could be forgotten in a couple of years. “There could be a disruptive technology at any point,” he concludes.
In November, I’m back in Houston to get my results. I’m about to learn whether there’s a genetic killer, or even a genetic pest, lurking within me.
While an Ion Torrent machine can sequence an exome in just a few hours, I found that the process slows down considerably when a hospital gets involved. For one thing, the machine’s software can identify gene variants of possible interest, but that raw data must then be scrutinized by experts. And my results had to be certified as medically accurate to meet the hospital’s stringent standards, which meant checking and double-checking. Lupski smiles across the table and says, “It’s a $1000 genome but a $100 000 analysis.”
These genetic results would ordinarily go to my physician, who would interpret them for me in light of my family medical history. The Baylor researchers don’t know that history, so they’ve pulled out everything that seems possibly significant. I’ll have to tell them if any of the findings match ailments that have afflicted my family members. “Okay, lay it on me,” I say.
The doctors hand me a six-page report with scary words that jump out: Parkinson’s, kidney failure, cardiomyopathy (a deterioration of the heart muscles). The doctors hurry to put these findings in context. “We did not find anything that requires immediate medical attention,” says Christine Eng, the physician and researcher who heads Baylor’s commercial genomics lab. That’s a relief. While I do have mutations in genes that are known to be associated with these life-ending maladies, my particular type of mutations haven’t been proved to cause a single tremor or irregular heartbeat.
Still, these findings are red flags, and one of those flags is waving in the breeze. My paternal grandfather’s kidneys failed toward the end of his life, I tell the doctors. Could he have had this same mutation?
“Aha,” Eng says, scribbling a note on her copy of the report. “This would be a situation where if we had more information from your family, we could rule this in or out as being significant for you.” If my doctor can get access to my grandfather’s medical records, she says, he can determine whether I’m likely to develop this particular kidney disease and come up with a monitoring plan. As I sit here with some of the nation’s top practitioners of genetic medicine, I imagine a future in which my exome data is integrated into my electronic medical records. At my first mention of kidney-related symptoms, a genetic alert for my doctor would pop up.
We go over the rest of the report, discussing a few recessive diseases that I seem to be a carrier for: There is something called Usher syndrome, for example, which typically causes congenital deafness. But I don’t know of any deafness in my family. And that’s about it. If the project shows the potential of personalized genetic medicine, it also demonstrates its current limitations. “There’s still a lot more that we have to learn,” admits Lupski. The crystal ball has turned up a lot of possibilities, and a few shadowy grim reapers are waving from the murky depths. But the results probably won’t change my life in any meaningful way.
When I tell my parents the results, I can’t help sounding disappointed. “It’s reassuring,” I say when we gather around the kitchen table. “But I expected to find out more that seemed relevant to our family.” My mother asks if I would recommend an exome scan to others, and I say no, not unless they’re trying to answer a specific medical question. It doesn’t make sense now for healthy people to run out and get their exomes sequenced, because you just don’t find out enough that’s “medically actionable,” as the Baylor doctors would put it. But the situation is changing incredibly fast. “Ten years down the line it might make sense,” I tell my mom.
Then comes a surprise that casts doubt on my first judgment and forces me to see exome sequencing in a new light. In the weeks following my meeting at Baylor I idly Google the various conditions listed in the report the doctors gave me. One afternoon I type in “Usher syndrome” and follow a link to a National Institutes of Health Web page about the disorder. A few sentences in, I feel a shock of recognition. The syndrome, I read, is associated not just with deafness but also with night blindness and severe balance problems. My mother has been completely unable to see in the dark for as long as she can remember, and both she and her older brother have gotten dangerously wobbly on their feet over the past decade.
I go back to talk to my mother again, and she gasps at my recitation of symptoms: “Oh my gosh, I’ve got that! And my brother too!” But as soon as she has internalized the news, she asks the inevitable question: Can anything be done about it? Not much, I tell her. She would have to undergo a genetic test of her own to see if she really does have a form of Usher’s, and even a definitive diagnosis wouldn’t lead to a treatment, because there are none. “You’re at the frontier of genetic medicine,” I say, partly because I don’t know what else to say. “It’s very rare that a patient can go to the doctor saying, ‘I know about my genes, and now you have to build your treatment plan around my genetic knowledge.’ ” She could be monitored for further vision loss, I offer.
I return to Guilford one more time to talk over my exome results with Rothberg. He flips the pages of my report with curiosity and listens as I explain that I’d hoped the scan would give me more prompts to take action. That’s what comes of being an early adopter, Rothberg says. “We’re at a tipping point,” he adds, “and we’re just starting to sequence individuals. So we don’t have a lot of information that correlates your sequence with outcome. Right now there’s only a handful of genes that we can tell you something about.”
But, he goes on, every day new patients get sequenced and researchers add fresh information to genetic databases. “That’s why it’s so important now to sequence tens of thousands of people and to keep track of their medical records, so we can annotate the rest of the genome,” he explains. “So that five years from now when somebody else looks at their genome they won’t get a little report. They’ll get a much bigger report with much more statistically significant information.”
Rothberg is sure that his technology will enable these massive genomic investigations that will give researchers a deeper understanding of our individual genetic destinies. But there will always be uncertainties—for most of us, the crystal ball will always be somewhat clouded. Every human being may be a unique assortment of As, Ts, Gs, and Cs, but our lives are also composed of choices: yes or no, smoking or nonsmoking, beef or tofu, stairs or elevator, live here or there, take this job or that. We’re all products of both our genetic vulnerabilities and our environments, which means we have at least some control over our fates.
On my first visit to Baylor, Lupski quoted a geneticist named Leena Peltonen-Palotie to me. “I always liked her saying that our genes are the paper and pen we are born with, but the story we will write is up to us, ” Lupski recalls. “Everybody makes the assumption that I’m a genetic determinist,” he says. “In fact, the more you practice, the more you realize that genetics are not the whole story. They’re one part of the story. We all have susceptibilities, and we can either ignore that information or maximize the utility of that information.”
In other words, I can use my exome scan to write a better story of my life. Starting now.
About the Author

Getting Personal With Gene Sequencing
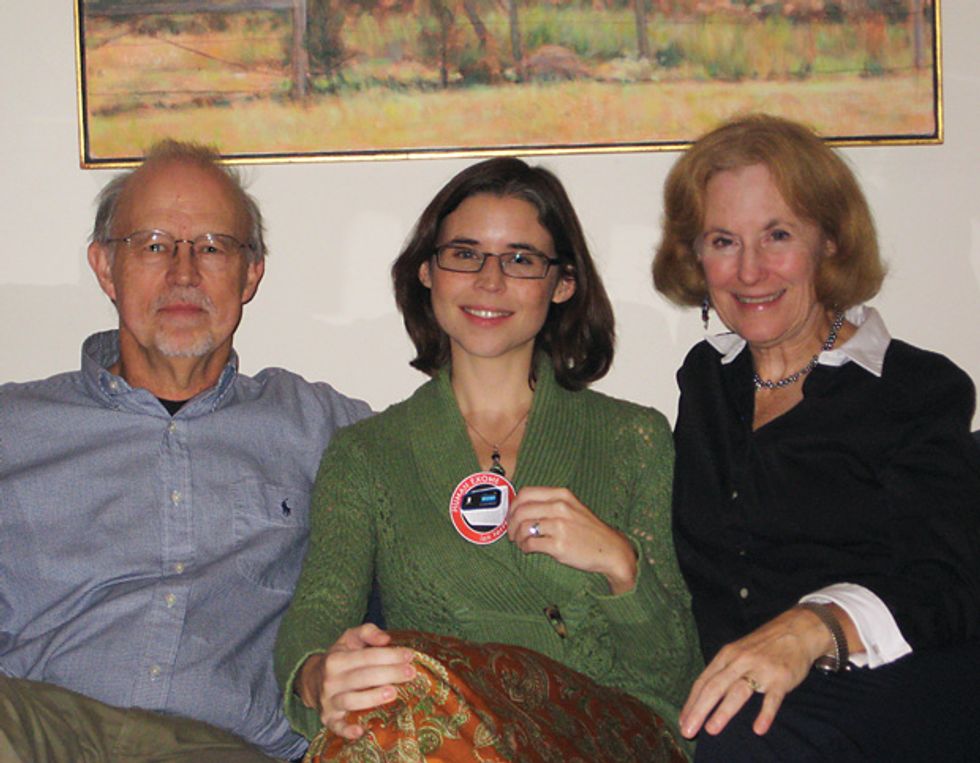
Associate Editor Eliza Strickland, who has been a science and technology journalist for most of her career, can make a personal case for genetic heritability: Her father is a research scientist, and her mother is a writer.
Of course, the genetic basis of personality and behavior is still mysterious, and researchers continue to debate questions of nature versus nurture. In medical science, though, we’re getting closer to real answers. Thanks to breathtaking advances in genetic sequencing, researchers are now finding genetic predispositions to such ailments as cancer and heart disease, and some physicians are offering genetic scans to patients to improve their treatments.
In “The Gene Machine and Me,” Strickland focuses on a new sequencing machine that will soon enable researchers to decode an entire human genome—a sequence of 3 billion molecules—for about US $1000. To make the story personal, Strickland had her own DNA run through the machine and had doctors analyze the resulting data. She asked the doctors to reveal everything they found in her genome, no matter how alarming, and she decided to share that information with IEEE Spectrum’s readers. That decision would affect her family: After all, she inherited her genetic material from her parents and shares it with her sister, niece, and nephew. Strickland describes the process of decoding her family’s genome both in these pages and in a one-hour radio special, which will be available on the Spectrum website and air on U.S. public radio stations this spring.
In the photo above, Strickland sits between her intrepid and obliging parents, holding a patch awarded to her by the researchers on completion of the sequencing project. We thank the Strickland family for baring all in the name of both science and journalism.
This article was modified on 14 March 2013.