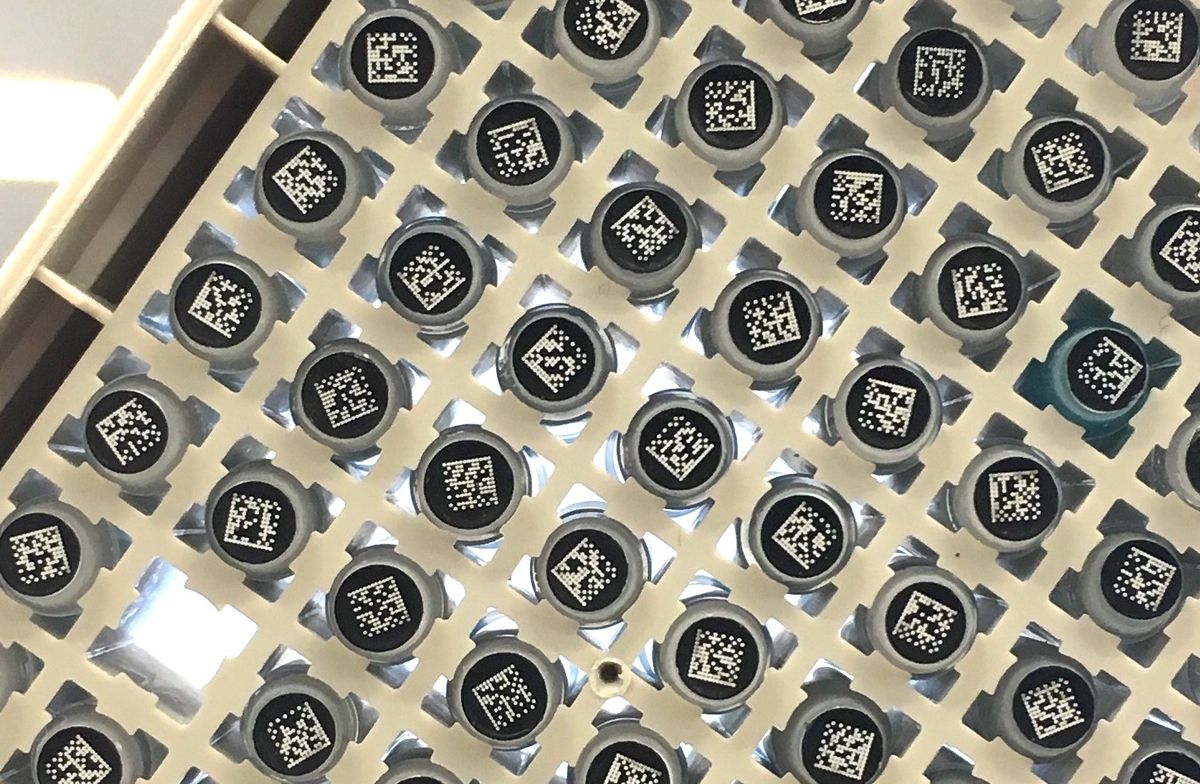
Try as a nefarious actor might, it would be near impossible to order the ingredients for making a deadly virus such as smallpox from scratch—at least not from any reputable company.
That’s because the world’s leading gene-synthesis firms all routinely screen customer requests against DNA sequences from hazardous viruses, bacteria, toxins and other ‘select agents.’ So, as soon as any would-be bioterrorist tried to purchase fragments of the pathogen’s genome, alarm bells would go off.
This biosecurity protocol is only as good as its database of potentially dangerous gene sequences, though. Case in point: Earlier this year, researchers from the University of Alberta demonstrated a potential vulnerability in the system when they reported the successful construction of horsepox virus, an extinct relative of smallpox, from DNA pieces ordered in the mail.
Horsepox is not on the list of verboten viruses, and the research itself (part of an effort to develop a new kind of vaccine) was in full compliance with government and university regulations. Still, the study sparked an uproar, and many in the biosecurity and biodefense communities are now openly questioning whether today’s screening tools are good enough to stop the threats posed by tomorrow’s synthetic bioweapons.
To help curtail those threats, the intelligence apparatus of the U.S. government is now looking to the world’s largest consumer of synthetic DNA for help. Last week, Boston-based Ginkgo Bioworks announced it had secured contracts worth up to $64 million to develop a range of biosecurity products for the nascent synthetic biology industry—chief among them, improved algorithms for screening orders made to gene-synthesis companies, and deep-learning models for detecting whether a DNA sample has been engineered in any way.
Todd Kuiken, a public policy researcher who studies synthetic biology at North Carolina State University, thinks Ginkgo is the right company for the job. “You need the experts developing the software,” he says, “and they are the best suited to figure this out.”
According to Patrick Boyle, head of design at Ginkgo, the company plans to take the algorithms it has developed over the past decade for identifying beneficial DNA sequences, and adapt the models for predicting whether bits of code could be potentially harmful.
Those algorithms have already led to the creation of new kinds of fragrance- and fertilizer-producing microbes. Now, with backing from the Intelligence Advanced Research Projects Agency, the company hopes to apply the tools to flag worrisome DNA sequence orders before they’re shipped and to discern whether an emerging pathogen circulating in nature evolved naturally or was engineered by humans.
These algorithmic and machine-learning approaches are what’s needed for biosecurity tools in the industry to move beyond simply matching sequences against known problematic strings of genetic code, says Laura Adam. Adam is cofounder of Ebiosec, a data management platform for synthetic biology, and the developer of GenoTHREAT, an open-source screening tool.
“The field is advancing, but no one is predicting things that are more novel,” she says. “A company like Ginkgo has real-life data that they can put into practice for detecting novel sequences.”
Boyle notes that Ginkgo already has the world’s largest database of engineered DNA sequences. “We’re now taking that database and training deep-learning models on that data to try to identify signatures of engineering that we may have unconsciously included in those designs.”
Ginkgo also has funding from the Department of Defense to develop enzymes that can sense and destroy chemical or biological threats. And the company is on retainer to further advance and commercialize these products and software tools if the initial research projects pan out.
Biosecurity tools may seem like a far cry from pharmaceuticals, fragrances and fertilizers. But, says Boyle, “We’re a platform company, so we’re really agnostic to our customer’s needs”—whether that customer is a crop company like Bayer, a drugmaker like Synlogic, or a branch of the U.S. military.
Plus, he adds, it’s imperative for Ginkgo to be a steward for responsible science if the billion-dollar biotech firm wants to realize the full potential for good within synthetic biology. “We just think it’s the right thing to do.”
Elie Dolgin is a science writer specializing in biomedical research and drug discovery. After a PhD spent studying the population genetics of nematodes, he swapped worms for words—entering journalism as an editor at The Scientist, Nature Medicine, and STAT. Now a freelancer, Elie is a frequent contributor to New Scientist, Nature, IEEE Spectrum, and more.