In cities around the world, an army of researchers has been busy swabbing mass transit kiosks, public benches, hospital floors and other high traffic urban surfaces. They’re looking for genetic evidence of SARS-CoV-2, the virus that causes COVID-19, and using the data to build computational models that could warn us of impending viral outbreaks.
“Instead of reactive epidemiology, it’s proactive tracing,” says Christopher Mason, a professor at Weill Cornell Medicine, who oversees the project. “If you did a simple targeted capture of known viruses once or twice a week in every city, you could catch an outbreak well before it becomes a big problem.”
The project, dubbed METACoV, spans 15 cities, including New York, Sao Paolo, Hong Kong, Tel Aviv and Berlin. It’s one of several such swabbing ventures by the international consortium Metagenomics and Metadesign of Subways and Urban Biomes, or MetaSUB. Since 2016, the organization has been collecting genetic samples from more than a hundred cities and studying what kind of microbial life exists on our most shared surfaces.
When the pandemic hit, MetaSUB’s global network got to work. The research groups send their postdocs, grad students, and medical students out into their cities to collect the samples. At public transit hubs, for example, they target features universal across nearly all cities: kiosks, benches, and turnstiles or other entry barriers. Then, for three awkward minutes, the researchers brush the surface with a nylon swab that looks like a Q-tip and is confirmed to be DNA and RNA free.
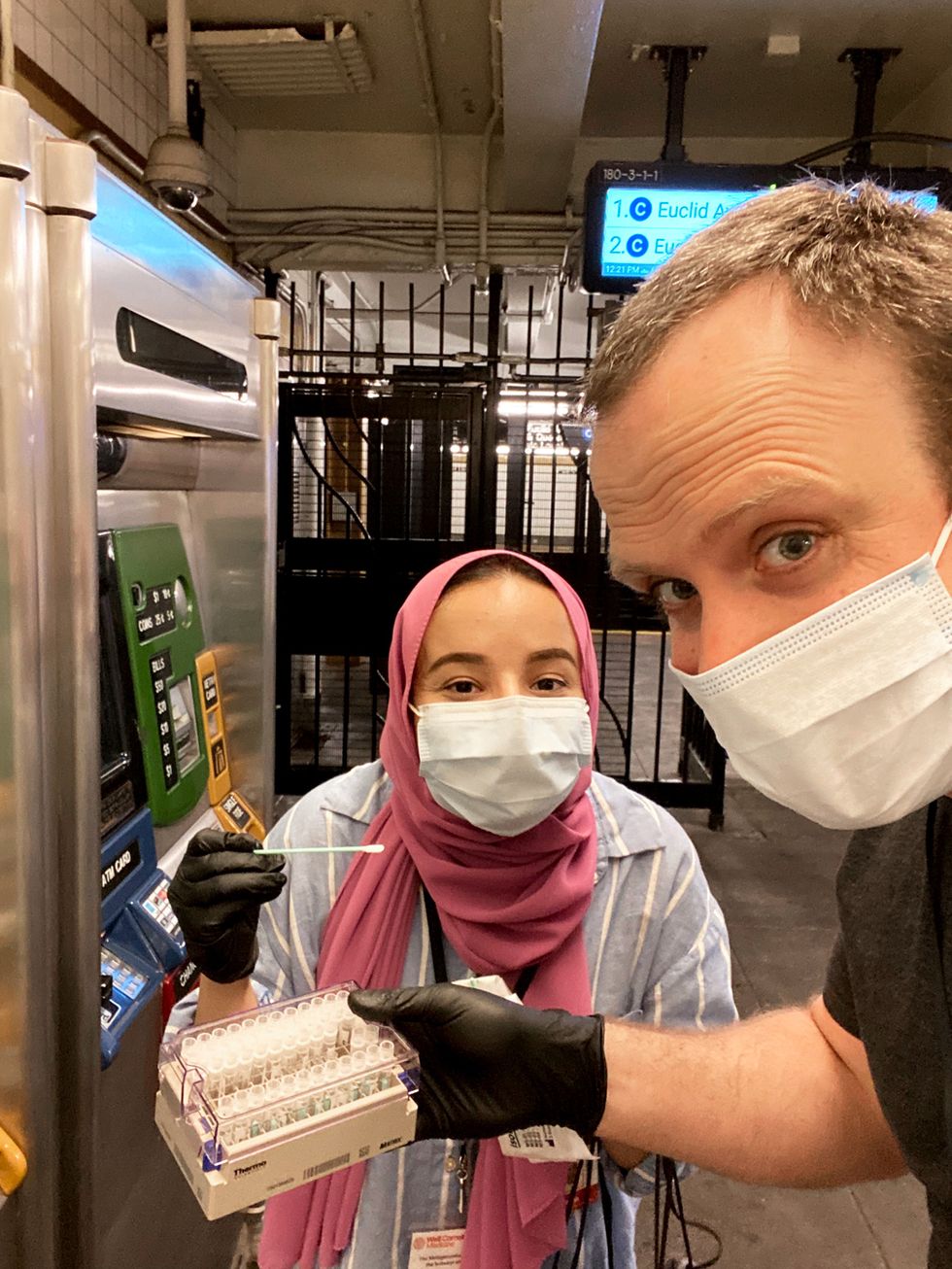

“Three minutes is the perfect balance between getting enough yield of DNA or RNA and minimizing the amount of social discomfort,” says Mason. “Pre-COVID, it always looked weird to be out there swabbing with gloves and a mask on, but the pandemic has really normalized our behavior.”
The researchers place the samples in a storage medium that preserves the genetic material, and then transport them back to the lab where the DNA or RNA is extracted, prepped, and sequenced. They use a range of bioinformatics tools to identify what viruses or species are present, and with what mutations.
The data enable researchers to create an atlas of sorts that shows where high risk variants of SARS-CoV-2 are emerging. This can help public health officials stay on top of the spread of the virus, and make decisions on where to focus vaccine supplies. Computational models also indicate where the virus might be headed genetically and geographically.
The data have yielded useful public health information. SARS-CoV-2 hasn’t shown up much on public transit surfaces, Mason says. But it is plentiful on hospital surfaces, including in rooms where people are negative for COVID-19, the researchers have found.
The team tested hospital samples to see how readily they could infect human cells. Luckily, none of the samples were successful. “It’s sitting on a hard surface that may have been bleached the night before. It’s a rough environment for anything to try to survive,” says Mason.
They also found SARS-CoV-2 on the ceilings of hospital restrooms. (If you’re eating while reading this, now would be a good time to put down your food.) How the virus reaches these ceilings is unclear, but Mason hypothesizes that it’s the result of what he calls “fecal fallout.” A COVID-19 positive person uses the toilet, flushes, and since the virus survives in stool and the toilets don’t have lids, the virus gets aerosolized and hits the ceiling. “It’s totally gross but probably true,” he says.
No wonder the consortium sends students to do the field work.
But it’s important work. In fact, stool may turn out to be a powerful predictor of outbreaks. As part of the METACoV project, the consortium is surveying city sewer systems, and what they are finding is fascinating: The amount of virus found in sewage typically peaks five to six days before confirmed cases peak in a population.
The National Institutes of Health (NIH) this year awarded a $5 million grant to Mason and his colleagues to support their wastewater surveillance project. The goal is develop an early warning and viral mapping system for genetic variants of SARS-CoV-2.
A paper summarizing the data collected so far from city centers is in press in Cell, Mason says, and data collected from hospitals is described in a paper published this month in the International Journal of Environmental Research and Public Health. The consortium continues to collect data from these environments. They are also studying the presence of the virus in mosquitoes and cats.
Mason’s team first published on the topic of urban metagenomics in 2015, when they analyzed the microbial ecosystem of the New York City subway system. Fun fact: They found that almost half of all DNA present on the subway’s surfaces matched no known organism.
Emily Waltz is a features editor at Spectrum covering power and energy. Prior to joining the staff in January 2024, Emily spent 18 years as a freelance journalist covering biotechnology, primarily for the Nature research journals and Spectrum. Her work has also appeared in Scientific American, Discover, Outside, and the New York Times. Emily has a master's degree from Columbia University Graduate School of Journalism and an undergraduate degree from Vanderbilt University. With every word she writes, Emily strives to say something true and useful. She posts on Twitter/X @EmWaltz and her portfolio can be found on her website.