Biotechnologists are jumping at the chance to use the revolutionary gene-editing tool known as CRISPR. The molecular gadget can be programmed to accurately tweak the DNA of any organism, but scientists need software algorithms to hasten the programming process. Dozens of teams are developing such software, and each faces the task of keeping up with rapidly evolving science and an increasingly crowded field.
CRISPR—short for Clustered, Regularly Interspaced, Short Palindromic Repeats—is a genetic phenomenon found in microbes that scientists adapted to disable a gene or add DNA at precise locations in the genetic code. CRISPR isn’t the first gene-editing tool on the block, but it is by far the simplest and cheapest, and since its adaptation four years ago, it has proliferated globally. Researchers can use it to knock out genes in animal models to study their function, give crops new agronomic traits, synthesize microbes that produce drugs, create gene therapies to treat disease, and potentially—after some serious ethical debate—to genetically correct heritable diseases in human embryos.
In less than four years, CRISPR “has transformed labs around the world,” says Jing-Ruey Joanna Yeh, a chemical biologist at Massachusetts General Hospital’s Cardiovascular Research Center, in Charlestown, who contributed to the development of the technology. “Because this system is so simple and efficient, any lab can do it.” Traditional genome modification techniques involve shuttling DNA into cells without knowing where in the genome it will stick. Editing with CRISPR is like placing a cursor between two letters in a word processing document and hitting “delete” or clicking “paste.” And the tool can cost less than US $50 to assemble. There are other genome-editing systems that are as precise as CRISPR, but they must be customized for every use and require far more expertise and resources to assemble.
As good as CRISPR is compared to its predecessors, the tool doesn’t always work, says Jacob Corn, scientific director at the Innovative Genomics Initiative at the University of California, Berkeley. “We don’t really understand why that is,” he says. That’s where software comes in. Algorithms can help researchers design their CRISPR tools in a way that is statistically more likely to succeed.
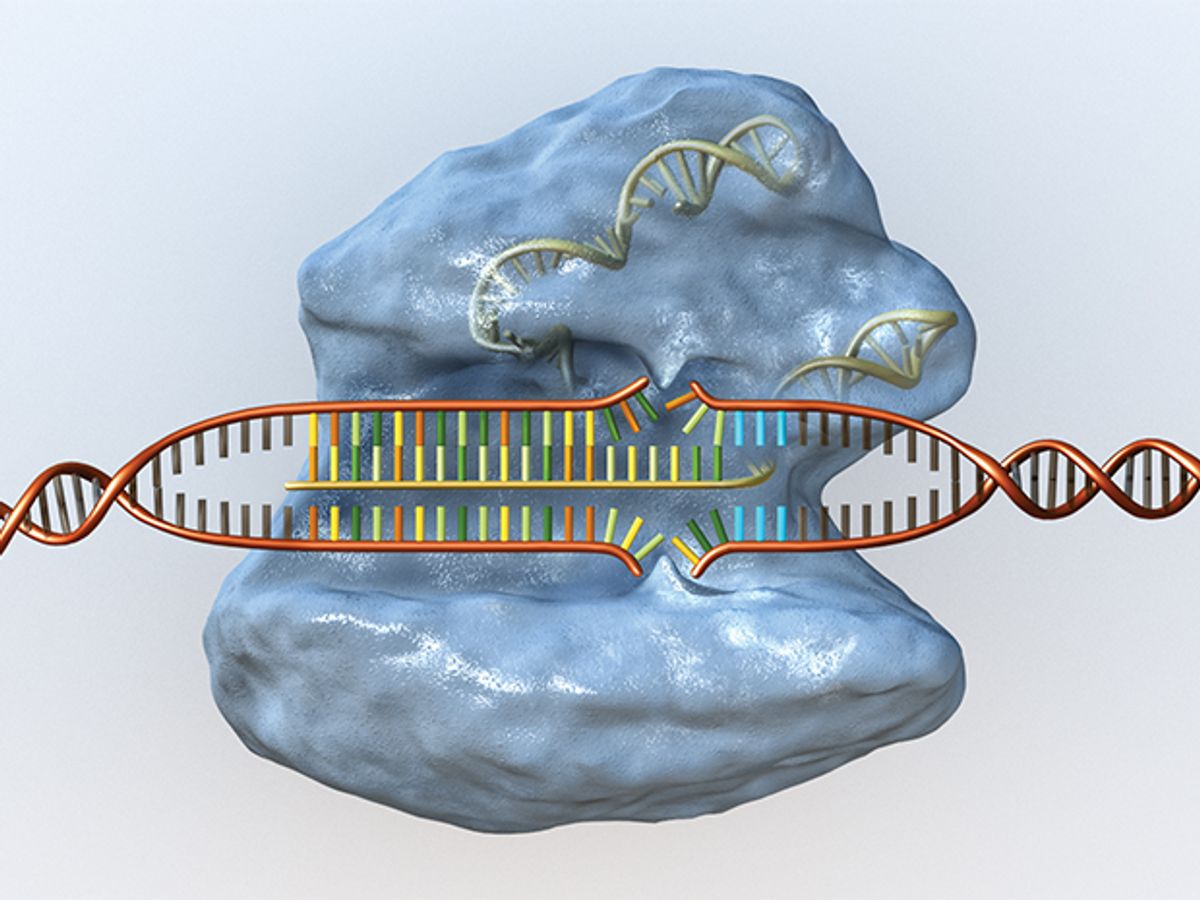
CRISPR systems are equipped with two main features: a short strand of programmable genetic code (called a guide RNA) and a protein (usually an enzyme called Cas9) that acts as a pair of molecular scissors. Once the complex is introduced into a cell, the guide RNA ushers Cas9 to a precise location in an organism’s DNA sequence (or genome), sticks to it like Velcro, and lets the Cas9 snip the DNA. The cell’s own machinery then repairs the cut, chewing up a bit of DNA or adding some in the process, thus disrupting the gene. Researchers can also intentionally introduce a piece of new genetic code to the site.
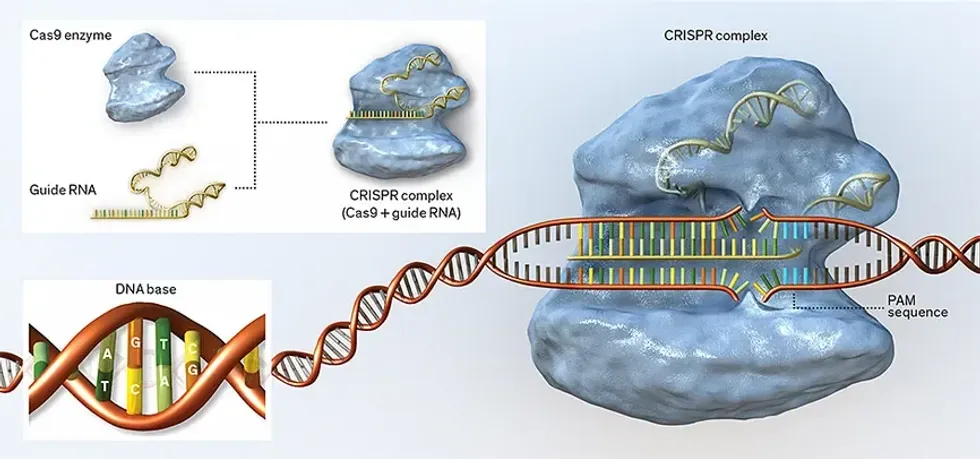
Guide RNAs find their targets in an organism’s genome by looking for a DNA segment with a complementary code of molecules. The molecules are called bases and are represented by the letters A (adenine), T (thymine), G (guanine), and C (cytosine).
DNA encodes genes as a sequence of chemicals (symbolized as A, T, G, and C) that mate with their chemical complements (A with T, G with C) to form the rungs of the molecular helix.Guide RNA ushers the CRISPR complex to a complementary site in the DNA, where the enzyme looks for landmarks called protospacer adjacent motifs, or PAMs. If the complex finds both a DNA match and a PAM, it snips the DNA strand so that either the gene’s sequence will be disrupted or a new piece of DNA can be inserted at the site.
Scientists editing the genome are usually looking for a segment that controls a particular function—a gene. Those are generally hundreds to thousands of bases long. Guide RNAs, however, are only about 20 bases long, so scientists must choose a complementary 20-base segment within the gene to target. There are two main constraints to consider: The target has to be located near a landmark that the molecular scissors can recognize, and it must be unlike any other 20-base segment anywhere else in the genome.
The Cas9 enzyme’s landmark is called a protospacer adjacent motif, or PAM. PAMs are easy to find in the genome—it’s like looking for the word “the” in a book, and any complementary 20-base segment adjacent to a PAM can work as the target site.
Ensuring that the 20-base segment is unique, however, is tougher. With a genetic code having only four letters and most organisms’ genomes ranging from millions to billions of base pairs, patterns are often repeated. Guide RNAs can get distracted by decoy segments, called off-target sites, and may end up mutating the wrong gene. Segments that differ from the target by just a couple of bases can trip up the tool. “You could scan across the whole genome by eye to try to find [off-targets], but it will take forever,” says Cameron Ross McPherson, a data scientist at the Institut Pasteur in Paris, who codeveloped the CRISPR software Protospacer Workbench.
Algorithms can do the search rapidly with few inputs from the user. Harvard University’s CHOPCHOP asks the user to enter the organism, the gene, and some optional advanced parameters. Within seconds, the algorithm finds within the target gene all the possible 20-base segments located near a PAM, ranks them based on their uniqueness in the genome and other parameters, and generates a list of guide RNAs to get you there. A search through the spaw gene in zebra fish, for example, yields 55 possible guide RNAs, most of which are unique sequences that differ from all other patterns in the genome by at least two bases.
Dozens of such software tools have popped up over the last two years, most of them available for free. A handful of companies, such as Benchling in San Francisco, offer more user-friendly interfaces than those of the free-to-the-public versions. But none of the software packages has emerged as a front-runner, says Michael Boutros at the German Cancer Research Center in Heidelberg, Germany, who codeveloped the CRISPR software E-CRISP.
Boutros says there is a lot of work to do. A list of 55 guide RNAs that theoretically might work is a helpful starting point, but it leaves researchers having to experiment, trial-and-error style, to see which one works the best. Algorithms that can instead predict with certainty that a particular guide RNA will work are needed.
To that end, biostatisticians are beginning to comb through experimental data to look for common patterns of successful guide RNAs that they can use to inform machine-learning-based prediction systems. But most of that data is scattered throughout small, individual studies. “Putting that all together will be a very powerful resource,” and it presents an opportunity for computer engineers to get involved, says UC Berkeley’s Corn. A few large data sets do exist already. A group from the Broad Institute in Cambridge, Mass., tested nearly 2,000 guide RNAs in human and mouse cells and recently published a set of rules for an improved algorithm.
Meanwhile, scientists are tinkering with Cas9 and other cutting proteins in an attempt to provide more options for CRISPR users. Some of those proteins improve the accuracy of the guide RNA. If they succeed, the need for software that predicts the accuracy of the tool may vanish, or at least evolve. “If we get to a place where we’ve absolutely, positively eliminated off-target effects, then great,” says Corn. “Are we there yet? No.”
This article appears in the March 2016 print issue as “Software Helps CRISPR Live Up to Its Hype.”