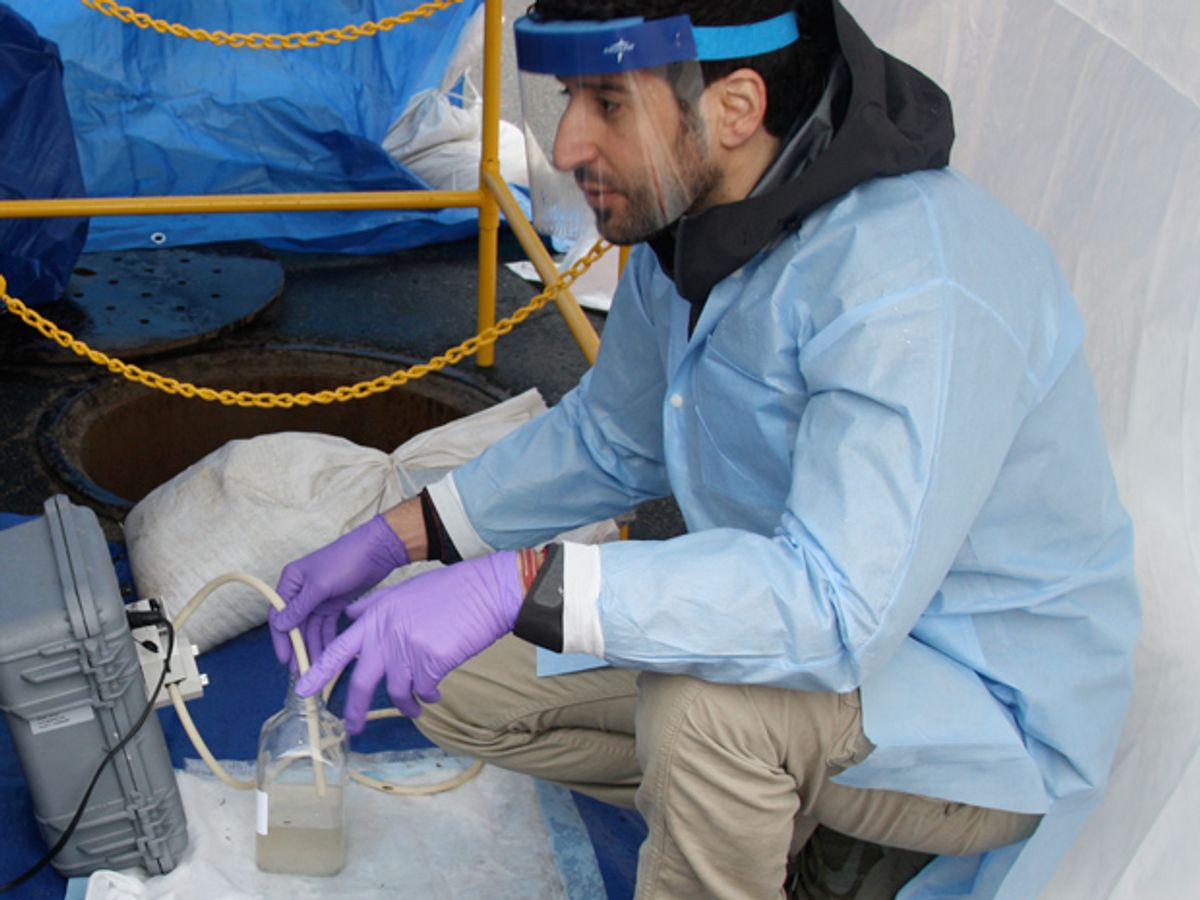
Ghaeli is an architect and research fellow at MIT who, beginning on Wednesday morning this week, oversaw a 24-hour effort to collect water samples from the sewer beneath an East Cambridge neighborhood. Every hour, the team pulled up half a liter of precious sludge.
The team was doing the groundwork for an ambitious project that aims to understand the wellbeing of a city by tracking its residents’ biological and chemical waste. Eventually, with robotic samplers placed below the streets, Cambridge may have a “smart sewer” that will let public health officials study the city’s collective microbiome—the communities of microorganisms that live in humans’ guts.
“The idea is to look at patterns of sewage relative to how we live our daily lives,” says Yaniv Jacob Turgeman, research director of the project. “We want this to be something that has actionable insights that enables public health in a meaningful way.”
The project—dubbed Underworlds, and supported by a US $4 million grant through the Kuwait-MIT Center for Natural Resources and the Environment—has been embraced by the city that the research team calls home. “If you get information closer to real time, you can rally resources to try to address a problem before it becomes a much bigger problem,” says Sam Lipson, director of environmental health for the City of Cambridge. “Those kinds of metrics are generally not available in public health.”
In the past, this type of “sewage epidemiology” has mostly been used to monitor population-level trends in illicit drug use. In Europe, for example, a recent continent-wide study of sewage from 42 cities revealed that cocaine and ecstasy use was greatest in large metropolises on weekends, whereas cannabis and methamphetamine use was more evenly distributed throughout the week in towns of all sizes.
The MIT team will also test for drugs—both illicit and pharmaceutical—but they plan to go much further. Led by Eric Alm, a computational microbiologist, and Carlo Ratti, an architect and engineer, the Underworlders will screen for viruses, such as influenza and norovirus, to detect incipient outbreaks in Cambridge. They will sequence the DNA of bacteria to identify food-borne pathogens. And they will search for biomarkers, or biochemical indicators of various aspects of human health and disease.
“The MIT project is extremely ambitious and pioneering,” says Christian Daughton, a chemist with the U.S. Environmental Protection Agency who spearheaded the idea of using sewage to track community health. “If this project proves successful in demonstrating some sort of proof of principal, it could represent a significant, seminal advancement in the prospects for quickly and inexpensively monitoring public health in real time.”
The researchers still have a number of logistical considerations to sort out. This week’s round-the-clock sampling should pinpoint the best times of day to pick up the strongest human signals. “We mainly want toilet water, as opposed to rain water or shower water or kitchen sink water,” explains Alm, co-director of the Center for Microbiome Informatics and Therapeutics at MIT. So before delving too deep into the mucky details of the project, he says, “we’re stepping back and saying: ‘How do we really nail down the collection protocols, and how do we really estimate what the demographics are so we can take it to that next level?’”
That next level will be far more automated. Ratti and his team at the MIT SENSEable City Lab are currently designing manhole-width, foot-tall robots that will be suspended on cables to move up and down in the sewer lines. Using a custom-made smartphone app, the researchers will be able to control the robots remotely to collect samples and feed data into a detailed sewage sampling information system. The robots will mostly serve as collection vehicles, although the researchers hope to embed sensors to measure temperature, flow, and other parameters.
The plan is to start testing the robot in the sewers in the coming months as the researchers ramp up to 10 sites throughout Cambridge. In this way, they hope to gain a sense of the geographic diversity of sewage signals across the city. Later this year, they will also begin to set up a similar platform in Kuwait, a country that’s almost 1,000-times the size of Cambridge. “The scale of what they’re trying to do is impressive,” says Ryan Newton, a microbial ecologist at the University of Wisconsin–Milwaukee who has worked with wastewater samples. “They should be able to get a really, really good handle on the variability in that population.”
More than one-third of Kuwaiti children are obese. Public health officials are particularly interested in learning whether sewage can help measure the impact of new policies designed to tackle expanding waistlines. Evidence for this idea comes from a study that Newton published in February in the journal mBio. Over the course of a year, Newton and his colleagues analyzed samples from wastewater treatment plants in 71 cities in 31 U.S. states. Because the composition of gut bacteria is intricately tied to health, Newton’s team could predict with close to 90 percent accuracy whether a microbial community came from a city whose residents were lean or obese. With enough long-term data from Kuwait, sewage analysis could theoretically reveal whether public health campaigns, such as a recent effort to curb the amount of sodium in bread, are having an impact on people’s metabolic wellbeing.
Similar efforts are gearing up around the world. As New York University biologist Jane Carlton will discuss at the upcoming Microbes In The City meeting in June, she and her colleagues are halfway through a two-year project to characterize the bacteria, viruses, and single-celled protists living in raw sewage flowing from all five boroughs of the Big Apple. Carlton is also in discussions about establishing a comparable sewer sampling initiative in Shanghai, China.
Meanwhile, a team from the Argonne National Laboratory (ANL) has begun a seven-year project to test for metabolites and microbial life from the sewer overflow pipes and rivers of Chicago. Like the Underworlds project, much of the data collection remains manual for now. But Jack Gilbert, a microbial ecologist who is leading the Chicago initiative, is working on a battery-powered sensor that runs DNA amplification reactions to search for any of 385 different organisms in a single sample of wastewater. The microfluidic device is currently housed in a box about the size of a suitcase and transmits data via Bluetooth.
“We hope to have these sensors in-line from the sewage water treatment plants,” says Gilbert. “The technology is still in a prototype form, but theoretically the microbiome and virulence factors associated with any potential threats could be automatically detected and relayed to a central system.”
The possibilities for the “sewage-ome” are near endless, says Alm. The challenge is to determine what’s feasible. “There are so many things you can do with the platform,” he says. “Let’s build it and then see what works.”
Elie Dolgin is a science writer specializing in biomedical research and drug discovery. After a PhD spent studying the population genetics of nematodes, he swapped worms for words—entering journalism as an editor at The Scientist, Nature Medicine, and STAT. Now a freelancer, Elie is a frequent contributor to New Scientist, Nature, IEEE Spectrum, and more.