As the deadly new coronavirus permeates the planet, scientists are using genetic sequencing and an open-source software tool to track its transmission.
The software tool, called Nextstrain, can’t predict where the virus is going next. But it can tell us where new cases of the virus are coming from. That’s crucial information for health officials globally, who are trying to determine whether new cases are arriving in their countries through international travel, or being transmitted locally.
This type of analysis, called genomic epidemiology, “is extremely valuable to public health,” says James Hadfield, a computational scientist working on Nextstrain. “The sooner we can turn around this data, the better the response can be.”
The novel coronavirus, which causes the respiratory disease COVID-19, first emerged in December in China, where it has infected over 80,000 people. It has since spread to more than 85 countries [PDF], with the largest concentrations of cases so far in South Korea, Iran, and Italy. More than 250 cases had been confirmed in the United States at press time.
As new cases pop up, it’s important to determine the virus’s origin—whether the infected person contracted the virus locally or in another region. That can inform decisions on travel restrictions, school closures, quarantines, and where to focus resources to contain the outbreak.
How genetic sequencing maps the coronavirus
Genomic analysis can provide clues about a virus’s origin. During an outbreak, a virus’s genetic code will steadily mutate as it spreads through a population. The mutations are slight—often just a single letter change in the code, like from AATC to ATTC. The mutations provide a time and geographical stamp of sorts. (There is no evidence that the mutations thus far affect the biology of the virus.)
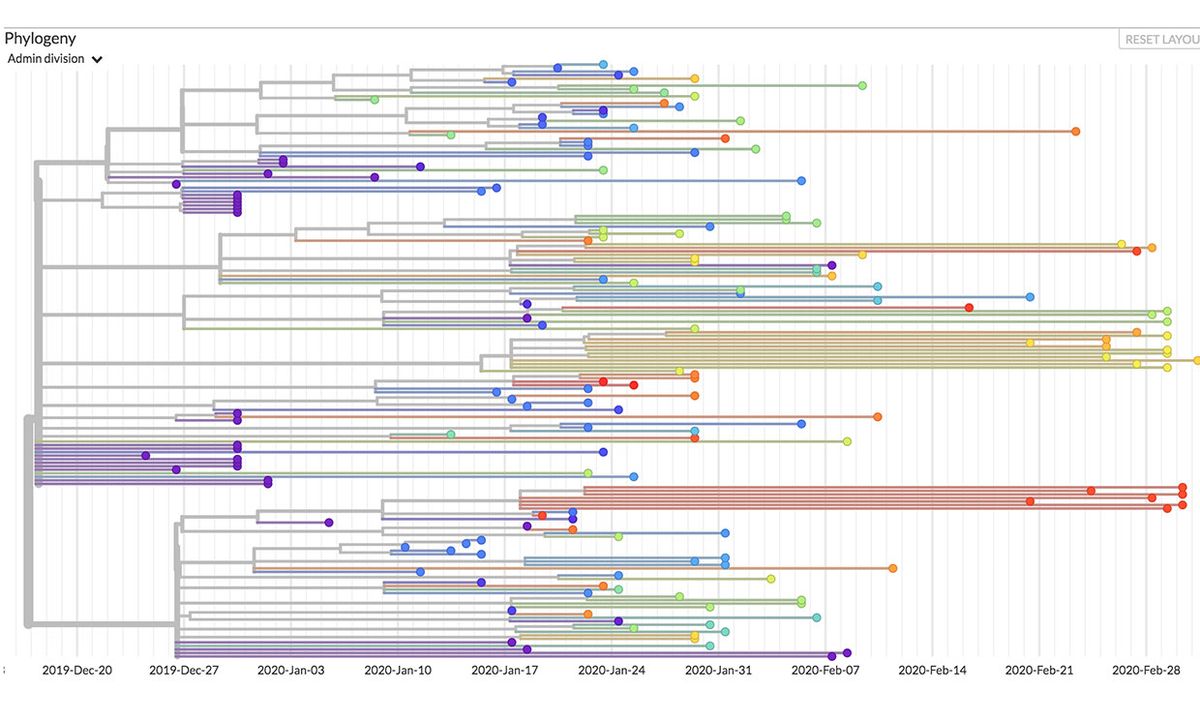
By comparing the genetic codes of viral samples taken globally, it’s possible to construct a map of the virus’s mutations as it moves around the world. That’s what Nextstrain does. “We rely on the presence of these naturally occurring genetic mutations to inform our visualizations of the virus’s spread,” says Hadfield.
Nextstrain charts a virus like a family tree, or evolutionary timeline. For coronavirus, that family tree originates in the Chinese city of Wuhan, and branches out from there. When new cases pop up, the genetic code of those viral samples can be compared to those in the database to determine its region of origin.
For example, in the United States, researchers have read, or sequenced, coronavirus genomes from eight cases in California. Of those, at least six were genetically distinct from each other, suggesting that they had all hitched rides to the United States through international travel, says Hadfield.
“What we can say from the genomic data is that there have been what looks like at least six independent introductions of the virus into California,” Hadfield says. “That’s not to say that ongoing local transmission in California is not occurring, but that the genomic data has not yet confirmed that.”
Coronavirus seeded in Seattle area
By contrast, the Seattle region has become a site of community transmission, according to Nextstrain’s analysis. The software compared two cases, one sampled in mid-January and the other sampled in late February, both in Snohomish County, near Seattle. The viruses were found to be genetically similar, suggesting local transmission.
Trevor Bedford, an investigator at the Fred Hutchinson Cancer Research Center, who co-developed Nextstrain, says that in the six weeks between the first and second cases, undetected community transmission was likely flourishing. He estimates that about 600 people in the Seattle area are likely infected, and possibly as many as 1,500. (Many cases of the novel coronavirus are mild, and infected people may not seek medical treatment.)
The Nextstrain endeavor relies, of course, on scientists being willing to obtain and sequence viral samples and upload them to freely accessible websites. And so far, researchers globally seem willing. Most are uploading sequencing data into the publicly available repository GISAID, says Hadfield. That’s where the Nextstrain team accesses its data, he says.
Scientists in resource-limited areas may not have the laboratory tools or training they need to perform this type of analysis. So a group called ARTIC Network has been providing protocols and training to enable scientists globally to perform disease surveillance and sequencing. They’re also developing a “lab-in-a-suitcase” that can be deployed to remote and resource-limited locations.
One success story came out of Brazil last week. In fewer than 48 hours, researchers collected a sample from an individual in São Paulo with coronavirus, sequenced the genome of the virus using ARTIC protocols, and shared the data on GISAID. The scientists used a portable genome sequencer called MinION.
Emily Waltz is a features editor at Spectrum covering power and energy. Prior to joining the staff in January 2024, Emily spent 18 years as a freelance journalist covering biotechnology, primarily for the Nature research journals and Spectrum. Her work has also appeared in Scientific American, Discover, Outside, and the New York Times. Emily has a master's degree from Columbia University Graduate School of Journalism and an undergraduate degree from Vanderbilt University. With every word she writes, Emily strives to say something true and useful. She posts on Twitter/X @EmWaltz and her portfolio can be found on her website.