Exabytes in a Test Tube: The Case for DNA Data Storage
With the right coding, the double helix could archive our entire civilization
Five thousand years ago, a man died in the Alps. It’s possible he died from a blow to the head, or he may have bled to death after being shot in the shoulder with an arrow. There’s a lot we don’t know about Ötzi (named for the Ötztal Alps, where he was discovered), despite the fact that researchers have spent almost 30 years studying him.
On the other hand, we know rather a lot about Ötzi’s physiological traits and even his clothes. We know he had brown eyes and a predisposition for cardiovascular diseases. He had type O positive blood and was lactose intolerant. The coat he was wearing was patched together using the leather of multiple sheep and goats, and his hat was made from a brown bear’s hide. All of this information came from sequencing the DNA of both Ötzi and the clothing he wore.
DNA can store remarkable amounts of genetic information and, as Ötzi demonstrates, can do so for thousands of years. The DNA molecule is a double-helix staircase of billions of molecular blocks, called base pairs, whose arrangement determines much of what makes each of us unique. Only recently have we contemplated using DNA to store electronic, digital data. And while DNA isn’t currently a viable alternative to memory sticks or hard-disk drives, it might be one of our best options to cope with the increasingly vast quantities of data we’ll create as data mining, analytics, and other big-data applications proliferate.
It was back in 2003 when some researchers, notably a group at the University of Arizona, became intrigued with the idea of using DNA to store data. But there were plenty of skeptics: Conventional mass-storage systems were doing the job cheaply and reliably. There was no compelling reason to seek out new options.
The situation has changed drastically over the last 15 years. We face an unprecedented data deluge in medicine, physics, astronomy, biology, and other sciences. The Sloan Digital Sky Survey, for example, produces about 73,000 gigabytes of data annually. At the European Organization for Nuclear Research (CERN), the Large Hadron Collider generates 50 million GB of data per year as it records the results of experiments involving, typically, 600 million particle collisions per second. These CERN results churn through a distributed computing grid comprising over 130,000 CPUs, 230 million GB of magnetic tape storage, and 300 million GB of online disk storage.
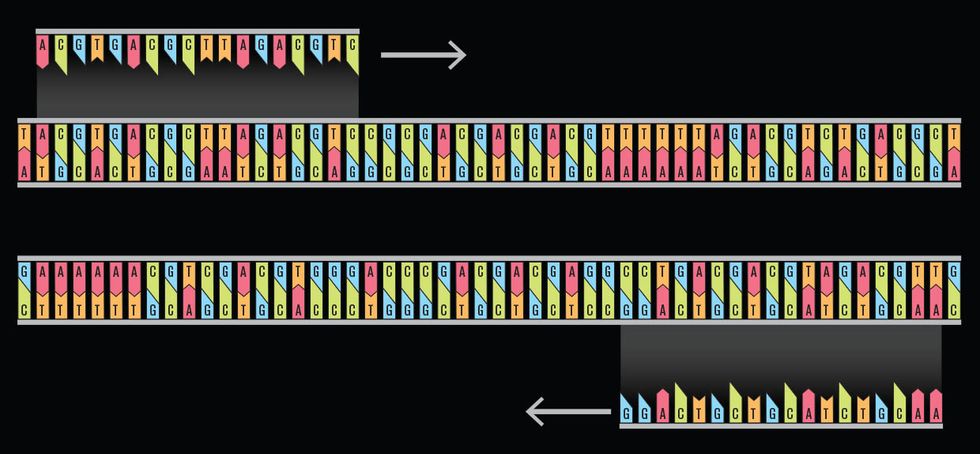
Using primers to replicate DNA
In the life sciences, DNA sequencing alone generates millions of gigabytes of data per year. Researchers predict that within a decade we will be swamped with 40 billion (109) GB of genomic data. All of that data will have to be stored for decades due to government regulations in the United States, Europe, and elsewhere.
Yet even as our data storage needs surge, traditional mass-storage technologies are starting to approach their limits. With hard-disk drives, we’re encountering a limit of 1 terabyte—1,000 GB—per square inch. Past that point, temperature fluctuations can induce the magnetically charged material of the disk to flip, corrupting the data it holds. We could try to use a more heat-resistant material, but we would have to drastically alter the technology we use to read and write on hard-disk drives, which would require huge new investments. The storage industry needs to look elsewhere.
DNA-based storage has come a long way since the early 2000s, when the technologies for reading DNA, let alone writing it, were still in their infancy. In those days, the Human Genome Project had only recently completed a draft of the human genome, at a mind-boggling cost exceeding US $2.7 billion, which works out to about $1 to read each base pair.
By the end of 2015, the cost for obtaining a highly accurate readout of an entire human genome had fallen below $1,500, according to the National Human Genome Research Institute. And today, roughly $1,000 is enough for you to get your entire genome sequenced. The cost of DNA sequencing is one three-millionth what it was 10 years ago.
Our ability to sequence, synthesize, and edit DNA has advanced at a previously inconceivable speed. Far from being expensive and impractical, these DNA technologies are the most disruptive in all of biotechnology. It’s now possible to write custom DNA strands for pennies per base pair, at least for short strands. Two companies, GenScript Biotech Corp. and Integrated DNA Technologies, provide DNA synthesis for 11 and 37 cents per base pair, respectively, for strands no longer than several hundred base pairs. Biotech startup companies buy their services and use the synthesized DNA to repair organs or create yeasts that produce unusual flavors to use in brewing beer.
For companies purchasing synthetic DNA, the cost depends on the length of the sequence being synthesized, because it is usually much more difficult to create long DNA strands. There are a handful of specialized efforts to synthesize longer strands—for example, an ongoing multilab effort is building an entirely synthetic yeast genome. Even so, commercially purchasing anything beyond 10,000 base pairs is currently impossible. (For reference, your genome has about 3.08 billion base pairs, a slightly smaller number than that of an African clawed frog.)
When reading DNA, sequencing devices produce fragments ranging in length from several hundred to tens of thousands of base pairs, which are then analyzed fragment by fragment before being stitched back together for a full readout. The whole process of reading an entire human genome takes less than a day. Researchers are now starting to sequence large quantities of fragments using nanopore technologies, which feed DNA through pores as if they were spaghetti noodles slipping through a large-holed strainer. As DNA passes through a pore, it can be read base by base.
In addition, DNA may be replicated exponentially at a low cost using the polymerase chain reaction, which duplicates a strand of DNA by splitting it apart and then building two identical strands by matching up the corresponding base pairs. These advances in reading DNA as well as in replicating it allow us, for the first time, to seriously consider DNA as a data-recording medium.
It still may not match other data storage options for cost, but DNA has advantages that other options can’t match. Not only is it easily replicated, it also has an ultrahigh storage density—as much as 100 trillion (1012) GB per gram. While the data representing a human genome, base pair by base pair, can be stored digitally on a CD with room to spare, a cell nucleus stores that same amount of data in a space about 1/24,000 as large. DNA does not have to be powered by an external energy source to retain data, as long as it’s stored in a controlled environment. And it can last for a long time: DNA can survive in less than ideal conditions for hundreds of thousands of years, although it often becomes highly degraded. After all, the Alps preserved Ötzi’s DNA for more than 5,000 years. Researchers once recovered DNA from the toe bones of a horse that had been preserved in a glacier for about 700,000 years.
Despite these appealing attributes, exploiting DNA for digital storage involves significant challenges. When it comes to building a storage system, the first task is to model the system’s structure and operation. To that end, two research groups—one at Harvard in 2012 and the other at the European Bioinformatics Institute, in the United Kingdom, in 2013—proposed conceptually simple designs for DNA-based storage.
Encoding text or binary code as DNA
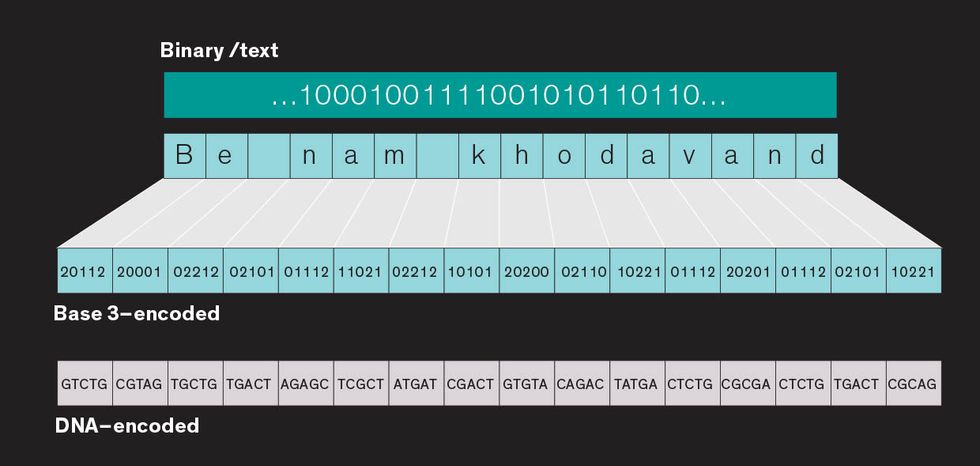

The basic idea was to convert the data into the DNA alphabet—adenine (A), thymine (T), cytosine (C), and guanine (G)—and store it in short strings with large amounts of overlap. The overlap would ensure that the data could be stitched back together accurately. For example, if the information was stored in strings that were 100 base pairs long, the last 75 base pairs from the previous string could be used as the first 75 base pairs for the next, with the next 25 base pairs tacked onto the end. With this strategy, the estimated cost of encoding 1 megabyte of data was over $12,000 for synthesizing the DNA and another $220 for retrieving it—rather prohibitively expensive at the moment.
-
Ensuring redundancy in DNA
Encoding data into a single long strand of DNA is asking for trouble when it comes time to recover the data. A safer process encodes the data in shorter strands. We then construct the first part of the next strand using the same data found at the end of the previous strand. This way we have multiple copies of the data for comparison.
-
Substitution errors in binary code
Damerau distance codes, which in natural-language processing are used to catch errors like misspellings (for example, “smort” instead of “smart”), can identify the spots in binary code where 1s and 0s have likely been substituted by mistake during copying or transcription.
-
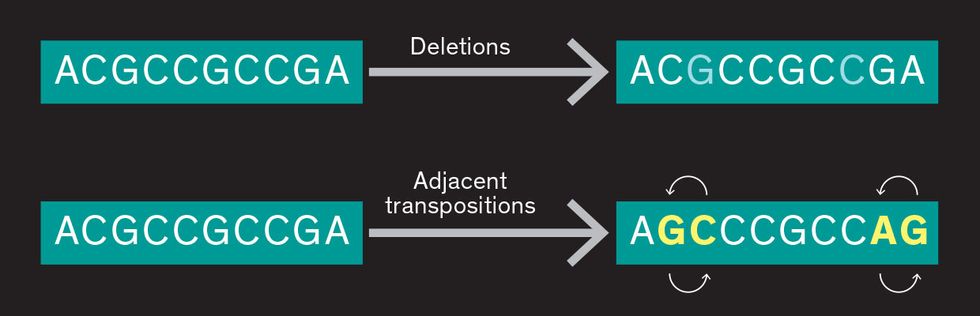
Substitution errors in DNA
Damerau distance codes can also be used to address the errors that occur in DNA, even though they’re more complex than binary errors. Sometimes bases are inadvertently deleted, and sometimes two will swap positions, errors that do not often occur in binary code.
Since then, research groups have demonstrated the long-term reliability of DNA-based data storage, the feasibility of using some traditional coding techniques, and even storing small amounts of data within the genomes of living bacteria. Our work, at the University of Illinois at Urbana-Champaign, in collaboration with the labs of Jian Ma and Huimin Zhao, pioneered random-access storage in DNA. We have been focused on solving the problems of random access, rewriting, and error-free data recovery for data that is read from DNA sequencing devices. Random access (the ability to directly access any information you want) and addressing (which tells you where to find that information) are key to any effective data storage method.
Our interest in DNA-based data storage emerged from our backgrounds in coding theory. Coding theory has made modern storage systems possible by enabling the proper data formatting for specific systems, the conversion of data from one format to another, and the correction of inevitable errors.
DNA-based storage systems are a tantalizing challenge for coding theorists. We were initially drawn to the challenge of identifying the sources of errors from both writing and reading DNA, and of developing coding techniques to correct or mitigate such errors. Coding improves the reliability of ultimately fallible storage devices and the feasibility of using cheaper options. But DNA-based storage systems are new and uncharted territory for coding theorists.
To understand the coding challenge presented by DNA, first consider a compact disc. The data is nicely organized into tracks, and we can easily access that data with the readily available hardware. DNA isn’t so simple. It’s inherently unordered; there are no tracks to follow to access the data.
A complete storage system would encompass many DNA molecules, so how would you even locate and select the specific molecule carrying the data you want? It would be like trying to fish a specific noodle out of a bowl of chicken noodle soup. It’s highly unlikely you’d grab the right noodle at random, but if you could replicate that specific noodle again and again, until you filled the bowl, any noodle you nab would likely be the right one.
Our idea for DNA data access is to synthesize each encoded strand with an additional sequence that acts as an address. Carefully designed sequences of the bases, called primers, would match that address sequence and begin the process of replicating the DNA of interest. In this way, we could exponentially reproduce DNA strands carrying the data of interest using the polymerase chain reaction, making it easy to find a copy of the right strand.
Of course, with DNA, it’s not quite so simple as plucking the right noodle out of your soup. Think of a primer as a sticky tape that binds to a specific set of rungs, or “complements,” on the DNA “ladder.” A primer should bind only to the specific address sequence it’s looking for. To make matters more difficult, not all primers are created equal: G and C base pairs typically bind more tightly than A and T, meaning that a primer constructed with too many A and T bases may not bind as strongly. Poorly designed primers can cause a lot of problems.
-
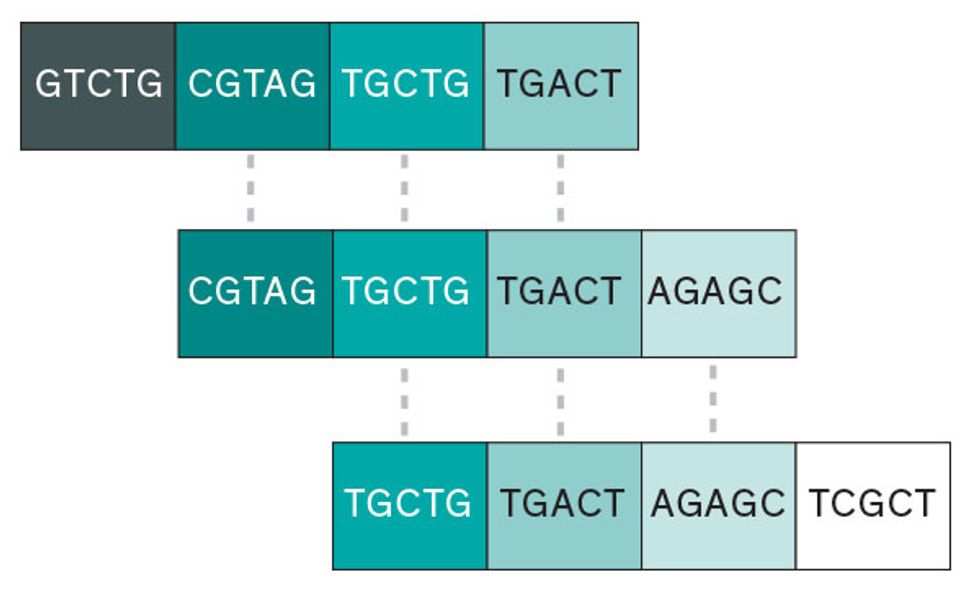
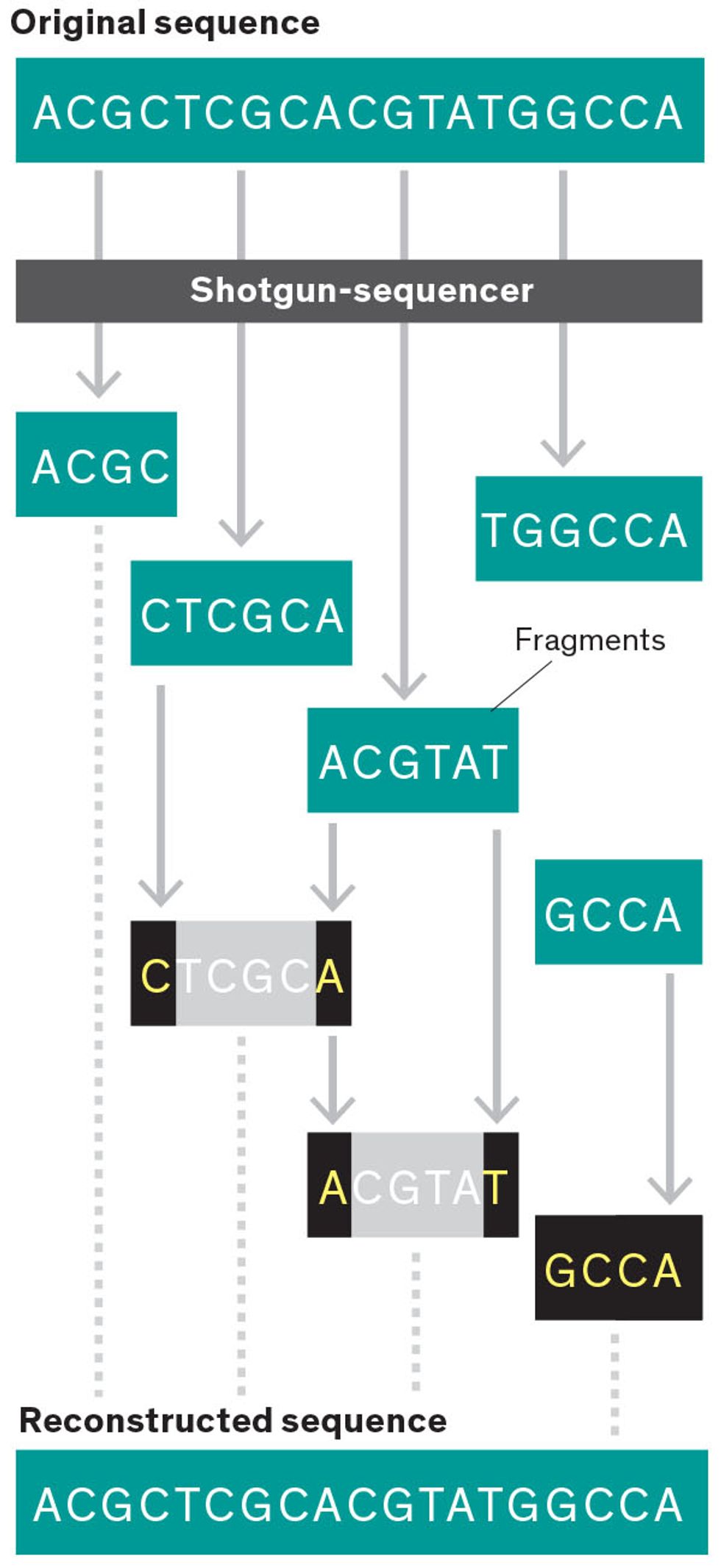
Reading DNA with a shotgun sequencer
“Shotgun-style” sequencing breaks copies of the long, unwieldy DNA strand into fragments of varying lengths. After those shorter segments are read, they can be compared with different fragments to reconstruct the entire sequence, although this method can introduce uncertainty about the placement of individual fragments.
-
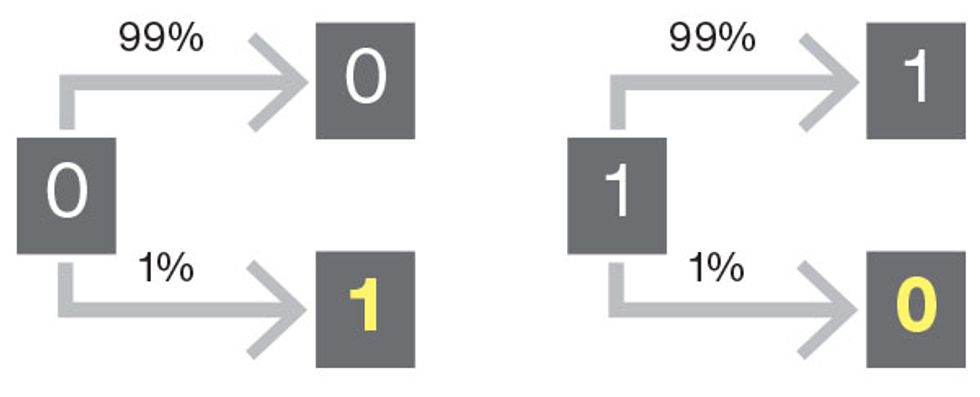
Transcription errors in binary code
When reading binary data from a traditional storage medium, there’s always a small chance that a 1 could be read as a 0 by mistake, or that a 0 could be read as a 1. Because we’re dealing with a simple two-state system, we can expect that each situation will occur with equal frequency.
We’re encountering intriguing coding questions in figuring out how to construct primers that will not only bind tightly but to the right targets. For example, because each primer will bind with its complement—A to T, G to C, and vice versa—how can we ensure that each address sequence doesn’t appear anywhere in the encoded data except as the address of the DNA strand you’re looking for? Otherwise, the primer may bind to the wrong location and replicate unwanted DNA.
Fortunately, coding theorists have been solving similar problems for traditional storage media for decades. Other challenges, for example, like those that emerge in connection with reading the DNA, aren’t typically encountered in conventional mass-storage systems. There are plenty of devices on the market that sequence DNA: Illumina’s HiSeq 2500 system, PacBio’s RS II and Sequel systems, and Oxford Nanopore Technologies’ MinION are just three examples. All such sequencers are prone to introducing different types of readout errors as they determine the exact sequence of As, Cs, Gs, and Ts that make up a DNA sample. Illumina devices, for example, sometimes substitute the wrong base when reading the strand—say, an A instead of a C. These errors become more frequent the further into the strand you get. The accidental deletion of entire blocks is also a concern, and nanopore sequencers often insert the wrong base pairs into readouts or omit base pairs entirely.
Different sequencers all require different code to compensate for their flaws. For Illumina sequencers, for example, we’ve proposed a coding scheme that adds redundancy to the sequence to eliminate the substitution errors that arise from the devices’ “shotgun-style” approach to sequencing. It’s tricky to rebuild a genome after breaking it apart to read individual sequences without occasionally inserting the wrong segment in the wrong location. Redundant sequences will improve the odds of recovering data even if a segment is corrupted as a result of being reassembled incorrectly.
For nanopore sequencers, we developed codes to address different types of substitution errors that arise from sequencing the strand too quickly. In traditional data storage, it’s just as likely that a 0 could be changed to a 1 as it is that a 1 could be changed to a 0. It’s not so simple with DNA, where an A could be rewritten as a T, C, or G, and the substitutions don’t happen with equal frequency. We’ve written codes to account for that fact, as well as codes to handle the base-pair deletions and swaps that naturally occur as DNA ages.
Nanopore-sequencer transcription errors in DNA
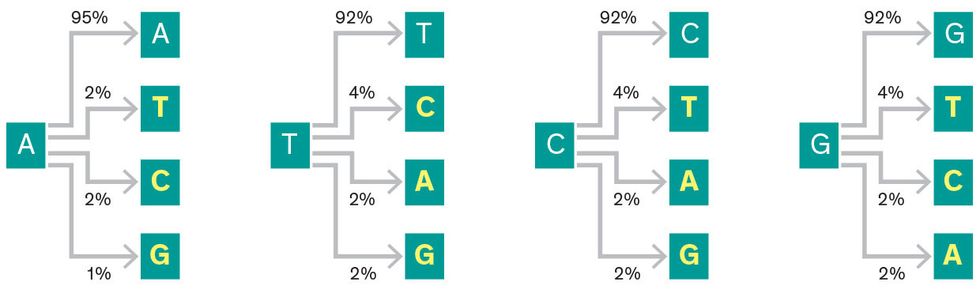
DNA-based storage, like any other data storage system, requires random access and efficient reading. But the biggest challenge is writing data inexpensively. Synthesizing DNA is still expensive, partly because of the molecule’s sheer complexity and partly because the market is not driving the development of cheaper methods. One possible approach to reduce costs is to prevent errors in the first place. By placing redundancies in the DNA sequences that store data, you can skip expensive after-the-fact corrections. This is common practice in every data storage method, but synthesizing companies currently aren’t equipped to pursue this—their production processes are so automated it would be prohibitively expensive to adjust them to produce these types of redundant strands.
Making DNA-based storage a practical reality will require cooperation among researchers on the frontiers of synthetic biology and coding theory. We’ve made big strides toward realizing a DNA-based storage system, but we need to develop systems to efficiently access the information encoded into DNA. We need to design coding schemes that guard against both synthesis and sequencing errors. And we need to figure out how to do these things cheaply.
If we can solve these problems, nature’s incredible storage medium—DNA—might also store our music, our literature, and our scientific advances. The very same medium that literally specifies who we are as individuals might also store our art, our culture, and our history as a species.
About the Authors
Olgica Milenkovic is a professor of electrical and computer engineering at the University of Illinois. Ryan Gabrys is a scientist with the U.S. Navy’s Spawar in San Diego and a postdoc at Illinois. Han Mao Kiah is a lecturer at Nanyang Technological University in Singapore and a former postdoc at Illinois. S.M. Hossein Tabatabaei Yazdi is a Ph.D. student working with Milenkovic.